Ubiquitination in T-Cell Activation and Checkpoint Inhibition: New Avenues for Targeted Cancer Immunotherapy
- PMID: 34639141
- PMCID: PMC8509743
- DOI: 10.3390/ijms221910800
Ubiquitination in T-Cell Activation and Checkpoint Inhibition: New Avenues for Targeted Cancer Immunotherapy
Abstract
The advent of T-cell-based immunotherapy has remarkably transformed cancer patient treatment. Despite their success, the currently approved immunotherapeutic protocols still encounter limitations, cause toxicity, and give disparate patient outcomes. Thus, a deeper understanding of the molecular mechanisms of T-cell activation and inhibition is much needed to rationally expand targets and possibilities to improve immunotherapies. Protein ubiquitination downstream of immune signaling pathways is essential to fine-tune virtually all immune responses, in particular, the positive and negative regulation of T-cell activation. Numerous studies have demonstrated that deregulation of ubiquitin-dependent pathways can significantly alter T-cell activation and enhance antitumor responses. Consequently, researchers in academia and industry are actively developing technologies to selectively exploit ubiquitin-related enzymes for cancer therapeutics. In this review, we discuss the molecular and functional roles of ubiquitination in key T-cell activation and checkpoint inhibitory pathways to highlight the vast possibilities that targeting ubiquitination offers for advancing T-cell-based immunotherapies.
Keywords: E3 ligases; T cells; cancer immunotherapy; checkpoint inhibition; deubiquitinases; ubiquitination.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

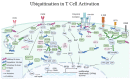

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

