Full-Length Transcriptome: A Reliable Alternative for Single-Cell RNA-Seq Analysis in the Spleen of Teleost Without Reference Genome
- PMID: 34646272
- PMCID: PMC8502891
- DOI: 10.3389/fimmu.2021.737332
Full-Length Transcriptome: A Reliable Alternative for Single-Cell RNA-Seq Analysis in the Spleen of Teleost Without Reference Genome
Abstract
Fish is considered as a supreme model for clarifying the evolution and regulatory mechanism of vertebrate immunity. However, the knowledge of distinct immune cell populations in fish is still limited, and further development of techniques advancing the identification of fish immune cell populations and their functions are required. Single cell RNA-seq (scRNA-seq) has provided a new approach for effective in-depth identification and characterization of cell subpopulations. Current approaches for scRNA-seq data analysis usually rely on comparison with a reference genome and hence are not suited for samples without any reference genome, which is currently very common in fish research. Here, we present an alternative, i.e. scRNA-seq data analysis with a full-length transcriptome as a reference, and evaluate this approach on samples from Epinephelus coioides-a teleost without any published genome. We show that it reconstructs well most of the present transcripts in the scRNA-seq data achieving a sensitivity equivalent to approaches relying on genome alignments of related species. Based on cell heterogeneity and known markers, we characterized four cell types: T cells, B cells, monocytes/macrophages (Mo/MΦ) and NCC (non-specific cytotoxic cells). Further analysis indicated the presence of two subsets of Mo/MΦ including M1 and M2 type, as well as four subsets in B cells, i.e. mature B cells, immature B cells, pre B cells and early-pre B cells. Our research will provide new clues for understanding biological characteristics, development and function of immune cell populations of teleost. Furthermore, our approach provides a reliable alternative for scRNA-seq data analysis in teleost for which no reference genome is currently available.
Keywords: full-length transcriptome; immune cell population; infection; scRNA-seq; teleost.
Copyright © 2021 Huang, Qiao, Xu, Gong, He, Qi, Gao, Cai, Grossart and Yan.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

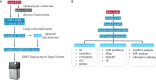






References
-
- Food and Agriculture Organization of the United Nations (FAO) . The State of World Fisheries and Aquaculture. Rome, Italy: FAO; (2016).
-
- Zhong Y, Qi W, Xu W, Zhao L, Xiao B, Yan Q, F. Insights Into Mesophilic Virulence, Antibiotic Resistant and Human Pathogenicity: A Genomics Study on the Aeromonas Salmonicida SRW-OG1 Newly Isolated From the Asian Fish Epinephelus Coioides. Aquaculture (2021) 539:736630. doi: 10.1016/j.aquaculture.2021.736630 - DOI
-
- He R, Zuo Y, Zhao L, Ma Y, Yan Q, Huang L. Copper Stress by Nutritional Immunity Activates the CusS-CusR Two-Component System That Contributes to Vibrio Alginolyticus Anti-Host Response But Affects Virulence-Related Properties. Aquaculture (2021) 532:736012. doi: 10.1016/j.aquaculture.2020.736012 - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

