Computer simulation of tumour resection-induced brain deformation by a meshless approach
- PMID: 34647427
- PMCID: PMC8881972
- DOI: 10.1002/cnm.3539
Computer simulation of tumour resection-induced brain deformation by a meshless approach
Abstract
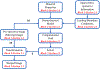
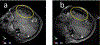
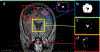
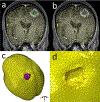
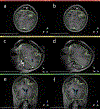
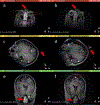
Tumour resection requires precise planning and navigation to maximise tumour removal while simultaneously protecting nearby healthy tissues. Neurosurgeons need to know the location of the remaining tumour after partial tumour removal before continuing with the resection. Our approach to the problem uses biomechanical modelling and computer simulation to compute the brain deformations after the tumour is resected. In this study, we use meshless Total Lagrangian explicit dynamics as the solver. The problem geometry is extracted from the patient-specific magnetic resonance imaging (MRI) data and includes the parenchyma, tumour, cerebrospinal fluid and skull. The appropriate non-linear material formulation is used. Loading is performed by imposing intra-operative conditions of gravity and reaction forces between the tumour and surrounding healthy parenchyma tissues. A finite frictionless sliding contact is enforced between the skull (rigid) and parenchyma. The meshless simulation results are compared to intra-operative MRI sections. We also calculate Hausdorff distances between the computed deformed surfaces (ventricles and tumour cavities) and surfaces observed intra-operatively. Over 80% of points on the ventricle surface and 95% of points on the tumour cavity surface were successfully registered (results within the limits of two times the original in-plane resolution of the intra-operative image). Computed results demonstrate the potential for our method in estimating the tissue deformation and tumour boundary during the resection.
Keywords: Total Lagrangian formulation; brain shift; medical image registration; meshless methods; patient-specific biomechanical simulations; tumour resection.
© 2021 John Wiley & Sons Ltd.
Figures







Similar articles
-
Beyond finite elements: a comprehensive, patient-specific neurosurgical simulation utilizing a meshless method.J Biomech. 2012 Oct 11;45(15):2698-701. doi: 10.1016/j.jbiomech.2012.07.031. Epub 2012 Aug 27. J Biomech. 2012. PMID: 22935689 Free PMC article.
-
SlicerCBM: automatic framework for biomechanical analysis of the brain.Int J Comput Assist Radiol Surg. 2023 Oct;18(10):1925-1940. doi: 10.1007/s11548-023-02881-7. Epub 2023 Apr 1. Int J Comput Assist Radiol Surg. 2023. PMID: 37004646 Free PMC article.
-
Suite of meshless algorithms for accurate computation of soft tissue deformation for surgical simulation.Med Image Anal. 2019 Aug;56:152-171. doi: 10.1016/j.media.2019.06.004. Epub 2019 Jun 12. Med Image Anal. 2019. PMID: 31229760 Free PMC article.
-
Biomechanical modeling and computer simulation of the brain during neurosurgery.Int J Numer Method Biomed Eng. 2019 Oct;35(10):e3250. doi: 10.1002/cnm.3250. Epub 2019 Sep 5. Int J Numer Method Biomed Eng. 2019. PMID: 31400252 Free PMC article. Review.
-
Breaking the state of the heart: meshless model for cardiac mechanics.Biomech Model Mechanobiol. 2019 Dec;18(6):1549-1561. doi: 10.1007/s10237-019-01175-9. Epub 2019 Jun 3. Biomech Model Mechanobiol. 2019. PMID: 31161351 Review.
Cited by
-
A position- and time-dependent pressure profile to model viscoelastic mechanical behavior of the brain tissue due to tumor growth.Comput Methods Biomech Biomed Engin. 2023 May;26(6):660-672. doi: 10.1080/10255842.2022.2082245. Epub 2022 May 31. Comput Methods Biomech Biomed Engin. 2023. PMID: 35638726 Free PMC article.
-
Automatic framework for patient-specific modelling of tumour resection-induced brain shift.Comput Biol Med. 2022 Apr;143:105271. doi: 10.1016/j.compbiomed.2022.105271. Epub 2022 Jan 30. Comput Biol Med. 2022. PMID: 35123136 Free PMC article.
-
Spatiotemporal single-cell tracking analysis in 3D tissues to reveal heterogeneous cellular response to mechanical stimuli.Sci Adv. 2023 Oct 13;9(41):eadf9917. doi: 10.1126/sciadv.adf9917. Epub 2023 Oct 13. Sci Adv. 2023. PMID: 37831766 Free PMC article.
References
-
- CANCER TODAY. Retrieved 16/12/2020, from World Health Organization. Global Health Observatory. Geneva: World Health Organization; https://gco.iarc.fr/today/home
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

