Emerging dynamics from high-resolution spatial numerical epidemics
- PMID: 34652271
- PMCID: PMC8568339
- DOI: 10.7554/eLife.71417
Emerging dynamics from high-resolution spatial numerical epidemics
Abstract
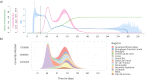
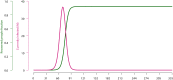
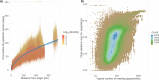
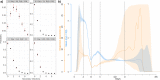
Simulating nationwide realistic individual movements with a detailed geographical structure can help optimise public health policies. However, existing tools have limited resolution or can only account for a limited number of agents. We introduce Epidemap, a new framework that can capture the daily movement of more than 60 million people in a country at a building-level resolution in a realistic and computationally efficient way. By applying it to the case of an infectious disease spreading in France, we uncover hitherto neglected effects, such as the emergence of two distinct peaks in the daily number of cases or the importance of local density in the timing of arrival of the epidemic. Finally, we show that the importance of super-spreading events strongly varies over time.
Keywords: computational biology; epidemiology; global health; high perfomance computing; parallel computing; systems biology.
© 2021, Thomine et al.
Conflict of interest statement
OT, SA, CB, MB, MS No competing interests declared
Figures

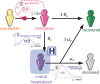
References
-
- Abar S, Theodoropoulos GK, Lemarinier P, O’Hare GMP. Agent based modelling and simulation tools: a review of the state-of-art software. Computer Science Review. 2017;24:13–33. doi: 10.1016/j.cosrev.2017.03.001. - DOI
-
- Aleta A, Martín-Corral D, Pastore Y Piontti A, Ajelli M, Litvinova M, Chinazzi M, Dean NE, Halloran ME, Longini IM, Merler S, Pentland A, Vespignani A, Moro E, Moreno Y. Modelling the impact of testing, contact tracing and household quarantine on second waves of COVID-19. Nature Human Behaviour. 2020;4:964–971. doi: 10.1038/s41562-020-0931-9. - DOI - PMC - PubMed
-
- Althaus CL, Turner KM, Schmid BV, Heijne JC, Kretzschmar M, Low N. Transmission of Chlamydia trachomatis through sexual partnerships: a comparison between three individual-based models and empirical data. Journal of The Royal Society Interface. 2012;9:136–146. doi: 10.1098/rsif.2011.0131. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

