Cell-free DNA from bile outperformed plasma as a potential alternative to tissue biopsy in biliary tract cancer
- PMID: 34653800
- PMCID: PMC8517551
- DOI: 10.1016/j.esmoop.2021.100275
Cell-free DNA from bile outperformed plasma as a potential alternative to tissue biopsy in biliary tract cancer
Abstract
Background: Biliary tract cancers (BTCs) are rare and highly heterogenous malignant neoplasms. Because obtaining BTC tissues is challenging, the purpose of this study was to explore the potential roles of bile as a liquid biopsy medium in patients with BTC.
Patients and methods: Sixty-nine consecutive patients with suspected BTC were prospectively enrolled in this study. Capture-based targeted sequencing was performed on tumor tissues, whole blood cells, plasma, and bile samples using a large panel consisting of 520 cancer-related genes.
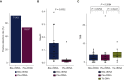
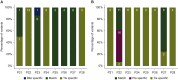
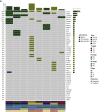
Results: Of the 28 patients enrolled in this cohort, tumor tissues were available in eight patients, and plasma and bile were available in 28 patients. Somatic mutations were detected in 100% (8/8), 71.4% (20/28), and 53.6% (15/28) of samples comprising tumor tissue DNA, bile cell-free DNA (cfDNA), and plasma cfDNA, respectively. Bile cfDNA showed a significantly higher maximum allele frequency than plasma cfDNA (P = 0.0032). There were 56.2% of somatic single-nucleotide variant (SNVs)/insertions and deletions (indels) shared between bile and plasma cfDNA. When considering the genetic profiles of tumor tissues as the gold standard, the by-variant sensitivity and positive predictive value for SNVs/indels in bile cfDNA positive for somatic mutations were both 95.5%. The overall concordance for SNVs/indels in bile was significantly higher than that in plasma (99.1% versus 78.3%, P < 0.0001). Moreover, the sensitivity of CA 19-9 combined with bile cfDNA achieved 96.4% in BTC diagnosis.
Conclusion: We demonstrated that bile cfDNA was superior to plasma cfDNA in the detection of tumor-related genomic alterations. Bile cfDNA as a minimally invasive liquid biopsy medium might be a supplemental approach to confirm BTC diagnosis.
Keywords: DNA; bile; biliary tract neoplasms; cell-free nucleic acids; liquid biopsy; sequence analysis.
Copyright © 2021 The Authors. Published by Elsevier Ltd.. All rights reserved.
Figures





Comment in
-
Bile cell‑free DNA: a game changer for management of biliary tract cancer?ESMO Open. 2022 Feb;7(1):100389. doi: 10.1016/j.esmoop.2022.100389. Epub 2022 Feb 2. ESMO Open. 2022. PMID: 35121524 Free PMC article. No abstract available.
References
-
- Lamarca A., Barriuso J., McNamara M.G., Valle J.W. Molecular targeted therapies: ready for “prime time” in biliary tract cancer. J Hepatol. 2020;73:170–185. - PubMed
-
- Valle J., Wasan H., Palmer D.H. Cisplatin plus gemcitabine versus gemcitabine for biliary tract cancer. N Engl J Med. 2010;362:1273–1281. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

