The translatome of neuronal cell bodies, dendrites, and axons
- PMID: 34670838
- PMCID: PMC8639352
- DOI: 10.1073/pnas.2113929118
The translatome of neuronal cell bodies, dendrites, and axons
Abstract
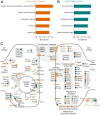
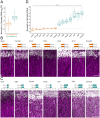
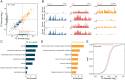
To form synaptic connections and store information, neurons continuously remodel their proteomes. The impressive length of dendrites and axons imposes logistical challenges to maintain synaptic proteins at locations remote from the transcription source (the nucleus). The discovery of thousands of messenger RNAs (mRNAs) near synapses suggested that neurons overcome distance and gain autonomy by producing proteins locally. It is not generally known, however, if, how, and when localized mRNAs are translated into protein. To investigate the translational landscape in neuronal subregions, we performed simultaneous RNA sequencing (RNA-seq) and ribosome sequencing (Ribo-seq) from microdissected rodent brain slices to identify and quantify the transcriptome and translatome in cell bodies (somata) as well as dendrites and axons (neuropil). Thousands of transcripts were differentially translated between somatic and synaptic regions, with many scaffold and signaling molecules displaying increased translation levels in the neuropil. Most translational changes between compartments could be accounted for by differences in RNA abundance. Pervasive translational regulation was observed in both somata and neuropil influenced by specific mRNA features (e.g., untranslated region [UTR] length, RNA-binding protein [RBP] motifs, and upstream open reading frames [uORFs]). For over 800 mRNAs, the dominant source of translation was the neuropil. We constructed a searchable and interactive database for exploring mRNA transcripts and their translation levels in the somata and neuropil [MPI Brain Research, The mRNA translation landscape in the synaptic neuropil. https://public.brain.mpg.de/dashapps/localseq/ Accessed 5 October 2021]. Overall, our findings emphasize the substantial contribution of local translation to maintaining synaptic protein levels and indicate that on-site translational control is an important mechanism to control synaptic strength.
Keywords: RNA localization; dendrites; local protein synthesis; translatome.
Copyright © 2021 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures


Comment in
-
A deep dive into local mRNA translation in neurons.Proc Natl Acad Sci U S A. 2021 Nov 9;118(45):e2117116118. doi: 10.1073/pnas.2117116118. Proc Natl Acad Sci U S A. 2021. PMID: 34737235 Free PMC article. No abstract available.
References
-
- Holt C. E., Martin K. C., Schuman E. M., Local translation in neurons: Visualization and function. Nat. Struct. Mol. Biol. 26, 557–566 (2019). - PubMed
-
- Hafner A. S., Donlin-Asp P. G., Leitch B., Herzog E., Schuman E. M., Local protein synthesis is a ubiquitous feature of neuronal pre- and postsynaptic compartments. Science 364, eaau3644 (2019). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

