Multi-Omics Analysis to Generate Hypotheses for Mild Health Problems in Monkeys
- PMID: 34677416
- PMCID: PMC8538200
- DOI: 10.3390/metabo11100701
Multi-Omics Analysis to Generate Hypotheses for Mild Health Problems in Monkeys
Abstract
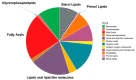
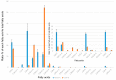
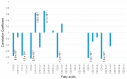
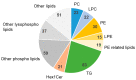
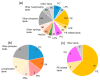
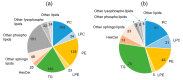
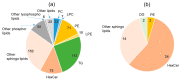
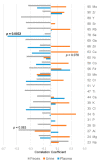
Certain symptoms associated with mild sickness and lethargy have not been categorized as definitive diseases. Confirming such symptoms in captive monkeys (Macaca fascicularis, known as cynomolgus monkeys) can be difficult; however, it is possible to observe and analyze their feces. In this study, we investigated the relationship between stool state and various omics data by considering objective and quantitative values of stool water content as a phenotype for analysis. By examining the food intake of the monkeys and assessing their stool, urine, and plasma, we attempted to obtain a comprehensive understanding of the health status of individual monkeys and correlate it with the stool condition. Our metabolomics data strongly suggested that many lipid-related metabolites were correlated with the stool water content. The lipidomic analysis revealed the involvement of saturated and oxidized fatty acids, metallomics revealed the contribution of selenium (a bio-essential trace element), and intestinal microbiota analysis revealed the association of several bacterial species with the stool water content. Based on our results, we hypothesize that the redox imbalance causes minor health problems. However, it is not possible to make a definite conclusion using multi-omics alone, and other hypotheses could be proposed.
Keywords: lipid mediators; lipidomics; metabolomics; metallomics; microbiota; multi-omics; multiple sample sources; redox; saturated fatty acids; selenium.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the interpretation of data or in the writing of the manuscript.
Figures

Similar articles
-
A Multi-omics Approach to Unraveling the Microbiome-Mediated Effects of Arabinoxylan Oligosaccharides in Overweight Humans.mSystems. 2019 May 28;4(4):e00209-19. doi: 10.1128/mSystems.00209-19. mSystems. 2019. PMID: 31138673 Free PMC article.
-
From Sample to Multi-Omics Conclusions in under 48 Hours.mSystems. 2016 Apr 26;1(2):e00038-16. doi: 10.1128/mSystems.00038-16. eCollection 2016 Mar-Apr. mSystems. 2016. PMID: 27822524 Free PMC article.
-
Stool microbiome and metabolome differences between colorectal cancer patients and healthy adults.PLoS One. 2013 Aug 6;8(8):e70803. doi: 10.1371/journal.pone.0070803. Print 2013. PLoS One. 2013. PMID: 23940645 Free PMC article.
-
Shotgun lipidomics in substantiating lipid peroxidation in redox biology: Methods and applications.Redox Biol. 2017 Aug;12:946-955. doi: 10.1016/j.redox.2017.04.030. Epub 2017 Apr 24. Redox Biol. 2017. PMID: 28494428 Free PMC article. Review.
-
Integration of lipidomics and metabolomics for in-depth understanding of cellular mechanism and disease progression.J Genet Genomics. 2020 Feb 20;47(2):69-83. doi: 10.1016/j.jgg.2019.11.009. Epub 2019 Dec 18. J Genet Genomics. 2020. PMID: 32178981 Review.
References
-
- Hird D.W., Anderson J.H., Bielitzki J.T. Diarrhea in nonhuman primates: A survey of primate colonies for incidence rates and clinical opinion. Lab. Anim. Sci. 1984;34:465–470. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

