Hybrid Sequencing in Different Types of Goat Skeletal Muscles Reveals Genes Regulating Muscle Development and Meat Quality
- PMID: 34679927
- PMCID: PMC8532877
- DOI: 10.3390/ani11102906
Hybrid Sequencing in Different Types of Goat Skeletal Muscles Reveals Genes Regulating Muscle Development and Meat Quality
Abstract
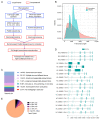
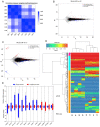
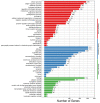
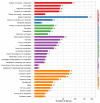
Domestic goats are commonly reared for meat and milk production in several regions of the world. However, the genetic mechanism underlying muscle development and meat quality of goats is limited. Therefore, the aim of this study was to identify known and novel genes regulating muscle development and meat quality of goats using second- and third-generation sequencing technologies. To achieve this, the meat quality and transcriptomes of longissimus dorsi (LD) and biceps femoris (BF) muscle tissues of Lingqiu Greyback goats were examined and compared. Differentially expressed genes (DEGs) and isoforms (DEIs) were functionally annotated. Results showed that 45,574 full-length transcripts covering 18,491 loci were characterized, and 12,566 genes were co-expressed in all samples. Differential expression analysis identified 231 DEGs, including 45 novel genes in the LD and BF muscles of the goats. Additionally, 1173 DEIs were found, in which 642 novel isoforms were identified in this study. Functional annotation and pathway analysis of the DEGs and DEIs revealed that some of them were associated with muscle growth and lipid metabolism. Overall, the findings of this study contribute to the understanding of the transcriptomic diversity underlying meat quality and muscle development of goat.
Keywords: full-length transcriptome; goat; growth; hybrid sequencing; meat quality; skeletal muscle.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures
Similar articles
-
Comparative Transcriptome Profile Analysis of Longissimus dorsi Muscle Tissues From Two Goat Breeds With Different Meat Production Performance Using RNA-Seq.Front Genet. 2021 Jan 13;11:619399. doi: 10.3389/fgene.2020.619399. eCollection 2020. Front Genet. 2021. PMID: 33519920 Free PMC article.
-
Comparative transcriptome and proteome analyses of the longissimus dorsi muscle for explaining the difference between donkey meat and other meats.Anim Biotechnol. 2023 Dec;34(7):3085-3098. doi: 10.1080/10495398.2022.2134883. Epub 2022 Oct 22. Anim Biotechnol. 2023. PMID: 36271875
-
Identification of differentially expressed genes through RNA sequencing in goats (Capra hircus) at different postnatal stages.PLoS One. 2017 Aug 11;12(8):e0182602. doi: 10.1371/journal.pone.0182602. eCollection 2017. PLoS One. 2017. PMID: 28800357 Free PMC article.
-
Effects of live weight at slaughter on fatty acid composition of Longissimus dorsi and Biceps femoris muscles of indigenous Lori goat.Trop Anim Health Prod. 2016 Jan;48(1):67-73. doi: 10.1007/s11250-015-0922-4. Epub 2015 Oct 2. Trop Anim Health Prod. 2016. PMID: 26431711
-
Candidate Genes and Their Expressions Involved in the Regulation of Milk and Meat Production and Quality in Goats (Capra hircus).Animals (Basel). 2022 Apr 11;12(8):988. doi: 10.3390/ani12080988. Animals (Basel). 2022. PMID: 35454235 Free PMC article. Review.
Cited by
-
Full-Length Transcriptome and Gene Expression Analysis of Different Ovis aries Adipose Tissues Reveals Transcript Variants Involved in Lipid Biosynthesis.Animals (Basel). 2023 Dec 19;14(1):7. doi: 10.3390/ani14010007. Animals (Basel). 2023. PMID: 38200738 Free PMC article.
-
Unraveling the molecular landscape of breast muscle development in domestic Yuzhong pigeons and European meat pigeon: Insights from Iso-seq and RNA-seq analysis.PLoS One. 2024 Jul 25;19(7):e0305907. doi: 10.1371/journal.pone.0305907. eCollection 2024. PLoS One. 2024. PMID: 39052586 Free PMC article.
-
Temporal Transcriptome Dynamics of Longissimus dorsi Reveals the Mechanism of the Differences in Muscle Development and IMF Deposition between Fuqing Goats and Nubian Goats.Animals (Basel). 2024 Jun 12;14(12):1770. doi: 10.3390/ani14121770. Animals (Basel). 2024. PMID: 38929389 Free PMC article.
-
Combined Analysis of Second- and Third-Generation Transcriptome Sequencing for Gene Characteristics and Identification of Key Splicing Variants in Wound Healing of Ganxi Goat Skin.Animals (Basel). 2024 Oct 26;14(21):3085. doi: 10.3390/ani14213085. Animals (Basel). 2024. PMID: 39518808 Free PMC article.
-
Integrative analysis of Iso-Seq and RNA-seq data reveals transcriptome complexity and differential isoform in skin tissues of different hair length Yak.BMC Genomics. 2024 May 21;25(1):498. doi: 10.1186/s12864-024-10345-8. BMC Genomics. 2024. PMID: 38773419 Free PMC article.
References
-
- Pophiwa P., Webb E.C., Frylinck L. A review of factors affecting goat meat quality and mitigating strategies. Small Rumin. Res. 2020;183:106035. doi: 10.1016/j.smallrumres.2019.106035. - DOI
-
- Shen J., Hao Z., Wang J., Hu J., Liu X., Li S., Ke N., Song Y., Lu Y., Hu L., et al. Comparative Transcriptome Profile Analysis of Longissimus dorsi Muscle Tissues from Two Goat Breeds with Different Meat Production Performance Using RNA-Seq. Front. Genet. 2020;11:619399. doi: 10.3389/fgene.2020.619399. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

