Genome wide association mapping for agronomic, fruit quality, and root architectural traits in tomato under organic farming conditions
- PMID: 34686145
- PMCID: PMC8532347
- DOI: 10.1186/s12870-021-03271-4
Genome wide association mapping for agronomic, fruit quality, and root architectural traits in tomato under organic farming conditions
Abstract
Background: Opportunity and challenges of the agriculture scenario of the next decades will face increasing demand for secure food through approaches able to minimize the input to cultivations. Large panels of tomato varieties represent a valuable resource of traits of interest under sustainable cultivation systems and for genome-wide association studies (GWAS). For mapping loci controlling the variation of agronomic, fruit quality, and root architecture traits, we used a heterogeneous set of 244 traditional and improved tomato accessions grown under organic field trials. Here we report comprehensive phenotyping and GWAS using over 37,300 SNPs obtained through double digest restriction-site associated DNA (dd-RADseq).
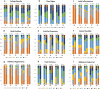
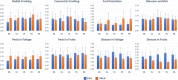
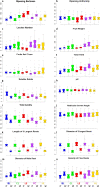
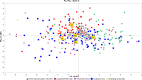
Results: A wide range of phenotypic diversity was observed in the studied collection, with highly significant differences encountered for most traits. A variable level of heritability was observed with values up to 69% for morphological traits while, among agronomic ones, fruit weight showed values above 80%. Genotype by environment analysis highlighted the strongest genotypic effect for aboveground traits compared to root architecture, suggesting that the hypogeal part of tomato plants has been a minor objective for breeding activities. GWAS was performed by a compressed mixed linear model leading to 59 significantly associated loci, allowing the identification of novel genes related to flower and fruit characteristics. Most genomic associations fell into the region surrounding SUN, OVATE, and MYB gene families. Six flower and fruit traits were associated with a single member of the SUN family (SLSUN31) on chromosome 11, in a region involved in the increase of fruit weight, locules number, and fruit fasciation. Furthermore, additional candidate genes for soluble solids content, fruit colour and shape were found near previously reported chromosomal regions, indicating the presence of synergic and multiple linked genes underlying the variation of these traits.
Conclusions: Results of this study give new hints on the genetic basis of traits in underexplored germplasm grown under organic conditions, providing a framework for the development of markers linked to candidate genes of interest to be used in genomics-assisted breeding in tomato, in particular under low-input and organic cultivation conditions.
Keywords: Genome-wide association mapping; Genotype by environment; Organic farming; Phenotyping; Tomato.
© 2021. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





References
-
- Anderson R, Bayer PE, Edwards D. Climate change and the need for agricultural adaptation. Curr Opin Plant Biol. 2020;56:197–202. - PubMed
-
- Le Campion A, Oury FX, Heumez E, Rolland B. Conventional versus organic farming systems: dissecting comparisons to improve cereal organic breeding strategies. Org Agr. 2020;10:63–74.
-
- The World of Organic Agriculture In: Statistics Sessions at BIOFACH 2020 Nürnberg, Germany. 2020. https://www.organic-world.net/yearbook/yearbook-2020.html. Accessed 20 June 2021.
-
- FAOSTAT 2019. http://www.fao.org/faostat/en/#home. Accessed 20 June 2021.
-
- Higashide T, Heuvelink E. Physiological and morphological changes over the past 50 years in yield components in tomato. J Am Soc Hortic Sci. 2009;134:460–465.

