Development and validation of a weakly supervised deep learning framework to predict the status of molecular pathways and key mutations in colorectal cancer from routine histology images: a retrospective study
- PMID: 34686474
- PMCID: PMC8609154
- DOI: 10.1016/S2589-7500(21)00180-1
Development and validation of a weakly supervised deep learning framework to predict the status of molecular pathways and key mutations in colorectal cancer from routine histology images: a retrospective study
Abstract
Background: Determining the status of molecular pathways and key mutations in colorectal cancer is crucial for optimal therapeutic decision making. We therefore aimed to develop a novel deep learning pipeline to predict the status of key molecular pathways and mutations from whole-slide images of haematoxylin and eosin-stained colorectal cancer slides as an alternative to current tests.
Methods: In this retrospective study, we used 502 diagnostic slides of primary colorectal tumours from 499 patients in The Cancer Genome Atlas colon and rectal cancer (TCGA-CRC-DX) cohort and developed a weakly supervised deep learning framework involving three separate convolutional neural network models. Whole-slide images were divided into equally sized tiles and model 1 (ResNet18) extracted tumour tiles from non-tumour tiles. These tumour tiles were inputted into model 2 (adapted ResNet34), trained by iterative draw and rank sampling to calculate a prediction score for each tile that represented the likelihood of a tile belonging to the molecular labels of high mutation density (vs low mutation density), microsatellite instability (vs microsatellite stability), chromosomal instability (vs genomic stability), CpG island methylator phenotype (CIMP)-high (vs CIMP-low), BRAFmut (vs BRAFWT), TP53mut (vs TP53WT), and KRASWT (vs KRASmut). These scores were used to identify the top-ranked titles from each slide, and model 3 (HoVer-Net) segmented and classified the different types of cell nuclei in these tiles. We calculated the area under the convex hull of the receiver operating characteristic curve (AUROC) as a model performance measure and compared our results with those of previously published methods.
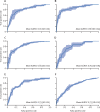
Findings: Our iterative draw and rank sampling method yielded mean AUROCs for the prediction of hypermutation (0·81 [SD 0·03] vs 0·71), microsatellite instability (0·86 [0·04] vs 0·74), chromosomal instability (0·83 [0·02] vs 0·73), BRAFmut (0·79 [0·01] vs 0·66), and TP53mut (0·73 [0·02] vs 0·64) in the TCGA-CRC-DX cohort that were higher than those from previously published methods, and an AUROC for KRASmut that was similar to previously reported methods (0·60 [SD 0·04] vs 0·60). Mean AUROC for predicting CIMP-high status was 0·79 (SD 0·05). We found high proportions of tumour-infiltrating lymphocytes and necrotic tumour cells to be associated with microsatellite instability, and high proportions of tumour-infiltrating lymphocytes and a low proportion of necrotic tumour cells to be associated with hypermutation.
Interpretation: After large-scale validation, our proposed algorithm for predicting clinically important mutations and molecular pathways, such as microsatellite instability, in colorectal cancer could be used to stratify patients for targeted therapies with potentially lower costs and quicker turnaround times than sequencing-based or immunohistochemistry-based approaches.
Funding: The UK Medical Research Council.
Copyright © 2021 The Author(s). Published by Elsevier Ltd. This is an Open Access article under the CC BY-NC-ND 4.0 license. Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
Declaration of interests MI reports grants from Roche, outside the submitted work. DS reports personal fees from Royal Philips, outside the submitted work. NMR reports research funding from GlaxoSmithKline and is also part of the PathLAKE consortium, which is partly funded by Royal Philips. All other authors declare no competing interests.
Figures


Comment in
-
Towards computationally efficient prediction of molecular signatures from routine histology images.Lancet Digit Health. 2021 Dec;3(12):e752-e753. doi: 10.1016/S2589-7500(21)00232-6. Epub 2021 Oct 19. Lancet Digit Health. 2021. PMID: 34686475 No abstract available.
References
-
- Al-Sohaily S, Biankin A, Leong R, Kohonen-Corish M, Warusavitarne J. Molecular pathways in colorectal cancer. J Gastroenterol Hepatol. 2012;27:1423–1431. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

