Quantification of marine benthic communities with metabarcoding
- PMID: 34687591
- PMCID: PMC9298412
- DOI: 10.1111/1755-0998.13536
Quantification of marine benthic communities with metabarcoding
Abstract
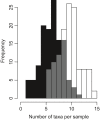
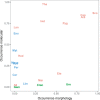
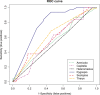
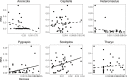
DNA metabarcoding methods have been implemented in studies aimed at detecting and quantifying marine benthic biodiversity. In such surveys, universal barcodes are amplified and sequenced from environmental DNA. To quantify biodiversity with DNA metabarcoding, a relation between the number of DNA sequences of a species and its biomass and/or the abundance is required. However, this relationship is complicated by many factors, and it is often unknown. In this study, we validate estimates of biomass and abundance from molecular approaches with those from the traditional morphological approach. Abundance and biomass were quantified from 126 samples of benthic intertidal mudflat using traditional morphological approaches and compared with frequency of occurrence and relative read abundance estimates from a molecular approach. A relationship between biomass and relative read abundance was found for two widely dispersed annelid taxa (Pygospio and Scoloplos). None of the other taxons, however, showed such a relationship. We discuss how quantification of abundance and biomass using molecular approaches are hampered by the ecology of DNA i.e. all the processes that determine the amount of DNA in the environment, including the ecology of the benthic species as well as the compositional nature of sequencing data.
Keywords: abundance; biomass; eDNA; metabarcoding; next-generation sequencing; quantification.
© 2021 The Authors. Molecular Ecology Resources published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Taxonomic accuracy and complementarity between bulk and eDNA metabarcoding provides an alternative to morphology for biological assessment of freshwater macroinvertebrates.Sci Total Environ. 2024 Jul 20;935:173243. doi: 10.1016/j.scitotenv.2024.173243. Epub 2024 May 16. Sci Total Environ. 2024. PMID: 38761946
-
Quantitative environmental DNA metabarcoding shows high potential as a novel approach to quantitatively assess fish community.Sci Rep. 2022 Dec 13;12(1):21524. doi: 10.1038/s41598-022-25274-3. Sci Rep. 2022. PMID: 36513686 Free PMC article.
-
Environmental DNA metabarcoding for benthic monitoring: A review of sediment sampling and DNA extraction methods.Sci Total Environ. 2022 Apr 20;818:151783. doi: 10.1016/j.scitotenv.2021.151783. Epub 2021 Nov 19. Sci Total Environ. 2022. PMID: 34801504 Review.
-
Environmental DNA metabarcoding: Transforming how we survey animal and plant communities.Mol Ecol. 2017 Nov;26(21):5872-5895. doi: 10.1111/mec.14350. Epub 2017 Oct 26. Mol Ecol. 2017. PMID: 28921802 Review.
-
Environmental DNA Metabarcoding: A Novel Method for Biodiversity Monitoring of Marine Fish Communities.Ann Rev Mar Sci. 2022 Jan 3;14:161-185. doi: 10.1146/annurev-marine-041421-082251. Epub 2021 Aug 5. Ann Rev Mar Sci. 2022. PMID: 34351788 Review.
Cited by
-
Monitoring insect biodiversity and comparison of sampling strategies using metabarcoding: A case study in the Yanshan Mountains, China.Ecol Evol. 2023 Apr 21;13(4):e10031. doi: 10.1002/ece3.10031. eCollection 2023 Apr. Ecol Evol. 2023. PMID: 37091562 Free PMC article.
-
SIBES: Long-term and large-scale monitoring of intertidal macrozoobenthos and sediment in the Dutch Wadden Sea.Sci Data. 2025 Feb 11;12(1):239. doi: 10.1038/s41597-025-04540-9. Sci Data. 2025. PMID: 39934128 Free PMC article.
-
High-throughput sequencing reveals omnivorous and preferential diets of the rotifer Polyarthra in situ.Front Microbiol. 2022 Dec 21;13:1048619. doi: 10.3389/fmicb.2022.1048619. eCollection 2022. Front Microbiol. 2022. PMID: 36620053 Free PMC article.
-
Advancing molecular macrobenthos biodiversity monitoring: a comparison between Oxford Nanopore and Illumina based metabarcoding and metagenomics.PeerJ. 2025 Apr 14;13:e19158. doi: 10.7717/peerj.19158. eCollection 2025. PeerJ. 2025. PMID: 40247828 Free PMC article.
References
-
- Alberdi, A. , Aizpurua, O. , Gilbert, M. T. P. , & Bohmann, K. (2018). Scrutinizing key steps for reliable metabarcoding of environmental samples. Methods in Ecology and Evolution, 9, 134–147. 10.1111/2041-210X.12849 - DOI
-
- Austen, M. C. , Lambshead, P. J. D. , Hutchings, P. A. , Boucher, G. , Heip, C. , King, G. , Koike, I. , Smith, C. , & Snelgrove, P. V. R. (2002). Biodiversity links above and below the marine sediment‐water interface that may influence community stability. Biodiversity and Conservation, 11, 113–136. 10.1023/A:1014098917535 - DOI
-
- Aylagas, E. , Borja, Á. , Muxika, I. , & Rodríguez‐Ezpeleta, N. (2018). Adapting metabarcoding‐based benthic biomonitoring into routine marine ecological status assessment networks. Ecological Indicators, 95, 194–202. 10.1016/j.ecolind.2018.07.044 - DOI
-
- Aylagas, E. , Mendibil, I. , Borja, Á. , & Rodríguez‐Ezpeleta, N. (2016). Marine sediment sample pre‐processing for macroinvertebrates metabarcoding: Mechanical enrichment and homogenization. Frontiers in Marine Science, 3, 1–8. 10.3389/fmars.2016.00203 - DOI
-
- Barnes, M. A. , & Turner, C. R. (2016). The ecology of environmental DNA and implications for conservation genetics. Conservation Genetics, 17, 1–17. 10.1007/s10592-015-0775-4 - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

