Comparison of Red-Complex Bacteria Between Saliva and Subgingival Plaque of Periodontitis Patients: A Systematic Review and Meta-Analysis
- PMID: 34692561
- PMCID: PMC8531218
- DOI: 10.3389/fcimb.2021.727732
Comparison of Red-Complex Bacteria Between Saliva and Subgingival Plaque of Periodontitis Patients: A Systematic Review and Meta-Analysis
Abstract
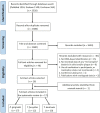
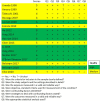
The development of periodontitis is associated with an imbalanced subgingival microbial community enriched with species such as the traditionally classified red-complex bacteria (Porphyromonas gingivalis, Tannerella forsythia, and Treponema denticola). Saliva has been suggested as an alternative to subgingival plaque for the microbial analysis due to its easy and non-invasive collection. This systematic review aims to determine whether the levels of red-complex bacteria assessed using saliva reflect those in subgingival plaque from periodontitis patients. The MEDLINE, EMBASE, and Cochrane Library databases were searched up to April 30, 2021. Studies were considered eligible if microbial data of at least one of the red-complex species were reported in both saliva and subgingival plaque from periodontitis patients, based on DNA-based methods. Of the 17 included studies, 4 studies used 16S rRNA gene sequencing techniques, and the rest used PCR-based approaches. The detection frequency of each red-complex species in periodontitis patients was reported to be > 60% in most studies, irrespective of samples types. Meta-analyses revealed that both detection frequencies and relative abundances of red-complex bacteria in saliva were significantly lower than those in subgingival plaque. Moreover, the relative abundances of all 3 bacterial species in saliva showed significantly positive correlation with those in subgingival plaque. In conclusion, current evidence suggests that one-time saliva sampling cannot replace subgingival plaque for microbial analysis of the red-complex bacteria in periodontitis patients. Given the positive microbial associations between saliva and subgingival plaque, a thorough review of longitudinal clinical studies is needed to further assess the role of saliva.
Keywords: 16S rRNA gene amplicon sequencing; Porphyromonas gingivalis; Tannerella forsythia; Treponema denticola; periodontitis; real-time PCR.
Copyright © 2021 Jiang, Song, Brandt, Cheng, Zhou, Exterkate, Crielaard and Deng.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures





Similar articles
-
Genetic analysis of Porphyromonas gingivalis (fimA), Aggregatibacter actinomycetemcomitans, and red complex in coronary plaque.J Investig Clin Dent. 2014 Aug;5(3):201-7. doi: 10.1111/jicd.12030. Epub 2013 Feb 27. J Investig Clin Dent. 2014. PMID: 23447375
-
Microbial analysis of subgingival plaque samples compared to that of whole saliva in patients with periodontitis.J Periodontol. 2014 Jun;85(6):819-28. doi: 10.1902/jop.2013.130306. Epub 2013 Oct 21. J Periodontol. 2014. PMID: 24144271
-
Oral microbiome in chinese patients with aggressive periodontitis and their family members.J Clin Periodontol. 2015 Nov;42(11):1015-23. doi: 10.1111/jcpe.12463. Epub 2015 Nov 14. J Clin Periodontol. 2015. PMID: 26412568
-
Influence of periodontal surgery on the subgingival microbiome-A systematic review and meta-analysis.J Periodontal Res. 2023 Apr;58(2):308-324. doi: 10.1111/jre.13092. Epub 2023 Jan 4. J Periodontal Res. 2023. PMID: 36597817
-
The Effect of Non-Surgical Periodontal Therapy on Subgingival Microbiota: A Systematic Review and Meta-Analysis.J Periodontal Res. 2025 May 9. doi: 10.1111/jre.13409. Online ahead of print. J Periodontal Res. 2025. PMID: 40347039 Review.
Cited by
-
Effects of Oleanolic Acid Derived from Wine Pomace on Periodontopathic Bacterial Growth in Healthy Individuals: A Randomized Placebo-Controlled Study.Dent J (Basel). 2024 May 8;12(5):133. doi: 10.3390/dj12050133. Dent J (Basel). 2024. PMID: 38786531 Free PMC article.
-
Are Salivary and Plasma Levels of Toll-Like Receptors 2 and 4 Elevated in Subjects With Chronic Periodontitis?: A Systematic Review and Meta-Analysis.Int J Inflam. 2025 May 6;2025:7405066. doi: 10.1155/ijin/7405066. eCollection 2025. Int J Inflam. 2025. PMID: 40371381 Free PMC article. Review.
-
Oral Spirochete Treponema denticola Intraoral Infection Reveals Unique miR-133a, miR-486, miR-126-3p, miR-126-5p miRNA Expression Kinetics during Periodontitis.Int J Mol Sci. 2023 Jul 28;24(15):12105. doi: 10.3390/ijms241512105. Int J Mol Sci. 2023. PMID: 37569480 Free PMC article.
-
Screening for Selenomonas noxia in a Pediatric and Adolescent Patient Population Reveals Differential Oral Prevalence across Age Groups.Int J Environ Res Public Health. 2024 Mar 23;21(4):391. doi: 10.3390/ijerph21040391. Int J Environ Res Public Health. 2024. PMID: 38673304 Free PMC article.
-
Salivary Inflammatory Mediator Profiles in Periodontal and Peri-Implant Health and Disease: A Cross-Sectional Study.Clin Implant Dent Relat Res. 2025 Feb;27(1):e70002. doi: 10.1111/cid.70002. Clin Implant Dent Relat Res. 2025. PMID: 39876538 Free PMC article.
References
-
- Belstrøm D., Sembler-Møller M. L., Grande M. A., Kirkby N., Cotton S. L., Paster B. J., et al. (2017). Microbial Profile Comparisons of Saliva, Pooled and Site-Specific Subgingival Samples in Periodontitis Patients. PloS One 12 (8), e0182992–e0182992. doi: 10.1371/journal.pone.0182992 - DOI - PMC - PubMed
-
- Boutaga K., Savelkoul P. H. M., Winkel E. G., van Winkelhoff A. J. (2007). Comparison of Subgingival Bacterial Sampling With Oral Lavage for Detection and Quantification of Periodontal Pathogens by Real-Time Polymerase Chain Reaction. J. Periodontol. 78 (1), 79–86. doi: 10.1902/jop.2007.060078 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

