Antibiotic-chemoattractants enhance neutrophil clearance of Staphylococcus aureus
- PMID: 34697316
- PMCID: PMC8546149
- DOI: 10.1038/s41467-021-26244-5
Antibiotic-chemoattractants enhance neutrophil clearance of Staphylococcus aureus
Abstract
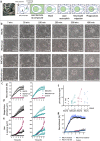
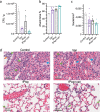
The pathogen Staphylococcus aureus can readily develop antibiotic resistance and evade the human immune system, which is associated with reduced levels of neutrophil recruitment. Here, we present a class of antibacterial peptides with potential to act both as antibiotics and as neutrophil chemoattractants. The compounds, which we term 'antibiotic-chemoattractants', consist of a formylated peptide (known to act as chemoattractant for neutrophil recruitment) that is covalently linked to the antibiotic vancomycin (known to bind to the bacterial cell wall). We use a combination of in vitro assays, cellular assays, infection-on-a-chip and in vivo mouse models to show that the compounds improve the recruitment, engulfment and killing of S. aureus by neutrophils. Furthermore, optimizing the formyl peptide sequence can enhance neutrophil activity through differential activation of formyl peptide receptors. Thus, we propose antibiotic-chemoattractants as an alternate approach for antibiotic development.
© 2021. The Author(s).
Conflict of interest statement
A.Y.P., J.A.E.P., J.T., and M.J.C. have filed for patent protection (PCT/AU2021/050306) of the antibiotic-chemoattractant compounds. The remaining authors declare no competing interests.
Figures



Similar articles
-
Differential effects of antibiotics on neutrophils exposed to lipoteichoic acid derived from Staphylococcus aureus.Ann Clin Microbiol Antimicrob. 2020 Nov 3;19(1):50. doi: 10.1186/s12941-020-00392-w. Ann Clin Microbiol Antimicrob. 2020. PMID: 33143710 Free PMC article.
-
Neutrophil chemotaxis by pathogen-associated molecular patterns--formylated peptides are crucial but not the sole neutrophil attractants produced by Staphylococcus aureus.Cell Microbiol. 2006 Feb;8(2):207-17. doi: 10.1111/j.1462-5822.2005.00610.x. Cell Microbiol. 2006. PMID: 16441432
-
Identification of formyl peptides from Listeria monocytogenes and Staphylococcus aureus as potent chemoattractants for mouse neutrophils.J Immunol. 2008 Jul 15;181(2):1429-37. doi: 10.4049/jimmunol.181.2.1429. J Immunol. 2008. PMID: 18606697 Free PMC article.
-
Neutrophil chemoattractant receptors in health and disease: double-edged swords.Cell Mol Immunol. 2020 May;17(5):433-450. doi: 10.1038/s41423-020-0412-0. Epub 2020 Apr 1. Cell Mol Immunol. 2020. PMID: 32238918 Free PMC article. Review.
-
Staphylococcus aureus versus neutrophil: Scrutiny of ancient combat.Microb Pathog. 2019 Jun;131:259-269. doi: 10.1016/j.micpath.2019.04.026. Epub 2019 Apr 16. Microb Pathog. 2019. PMID: 31002964 Review.
Cited by
-
Baohuoside I targets SaeR as an antivirulence strategy to disrupt MRSA biofilm formation and pathogenicity.NPJ Biofilms Microbiomes. 2025 Mar 19;11(1):45. doi: 10.1038/s41522-025-00681-2. NPJ Biofilms Microbiomes. 2025. PMID: 40102466 Free PMC article.
-
Discovery of phosphorylated lantibiotics with proimmune activity that regulate the oral microbiome.Proc Natl Acad Sci U S A. 2023 May 30;120(22):e2219392120. doi: 10.1073/pnas.2219392120. Epub 2023 May 22. Proc Natl Acad Sci U S A. 2023. PMID: 37216534 Free PMC article.
-
Vancomycin Resistance in Enterococcus and Staphylococcus aureus.Microorganisms. 2022 Dec 21;11(1):24. doi: 10.3390/microorganisms11010024. Microorganisms. 2022. PMID: 36677316 Free PMC article. Review.
-
Tranexamic acid protects against implant-associated infection by reducing biofilm formation.Sci Rep. 2022 Mar 22;12(1):4840. doi: 10.1038/s41598-022-08948-w. Sci Rep. 2022. PMID: 35318416 Free PMC article.
-
Immortalized mammosphere-derived epithelial cells retain a bioactive secretome with antimicrobial, regenerative, and immunomodulatory properties.Stem Cell Res Ther. 2024 Nov 14;15(1):429. doi: 10.1186/s13287-024-04019-1. Stem Cell Res Ther. 2024. PMID: 39543714 Free PMC article.
References
-
- CDC. Antibiotic resistance threats in the United States. (Report, 2013).
-
- O’Neill, J. The Review on antimicrobial resistance. (Report, 2016).
-
- Pfaller, M. A., Carvalhaes, C. G., Smith, C. J., Diekema, D. J. & Castanheira, M. Bacterial and fungal pathogens isolated from patients with bloodstream infection: frequency of occurrence and antimicrobial susceptibility patterns from the SENTRY Antimicrobial Surveillance Program (2012–2017). Diagn. Microbiol. Infect. Dis.97, 115016 (2020). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

