The solid and liquid states of chromatin
- PMID: 34717733
- PMCID: PMC8557566
- DOI: 10.1186/s13072-021-00424-5
The solid and liquid states of chromatin
Abstract
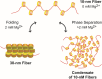
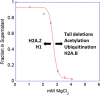
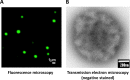
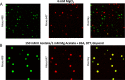
The review begins with a concise description of the principles of phase separation. This is followed by a comprehensive section on phase separation of chromatin, in which we recount the 60 years history of chromatin aggregation studies, discuss the evidence that chromatin aggregation intrinsically is a physiologically relevant liquid-solid phase separation (LSPS) process driven by chromatin self-interaction, and highlight the recent findings that under specific solution conditions chromatin can undergo liquid-liquid phase separation (LLPS) rather than LSPS. In the next section of the review, we discuss how certain chromatin-associated proteins undergo LLPS in vitro and in vivo. Some chromatin-binding proteins undergo LLPS in purified form in near-physiological ionic strength buffers while others will do so only in the presence of DNA, nucleosomes, or chromatin. The final section of the review evaluates the solid and liquid states of chromatin in the nucleus. While chromatin behaves as an immobile solid on the mesoscale, nucleosomes are mobile on the nanoscale. We discuss how this dual nature of chromatin, which fits well the concept of viscoelasticity, contributes to genome structure, emphasizing the dominant role of chromatin self-interaction.
© 2021. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

