High-Throughput On-Chip Human Mesenchymal Stromal Cell Potency Prediction
- PMID: 34725948
- PMCID: PMC8770576
- DOI: 10.1002/adhm.202101995
High-Throughput On-Chip Human Mesenchymal Stromal Cell Potency Prediction
Abstract
Human mesenchymal stromal cells (hMSCs) are a promising source for regenerative cell therapy. However, hMSC clinical use has been stymied by product variability across hMSC donors and manufacturing practices resulting in inconsistent clinical outcomes. The inability to predict hMSC clinical efficacy, or potency, is a major limitation for market penetration. Standard metrics of hMSC potency employ hMSCs and third-party immune cell co-cultures, however, these assays face translational challenges due to third-party donor variability and lack of scalability. While surrogate markers of hMSC potency have been suggested, none have yet had translational success. To address this, a high-throughput, scalable, low-cost, on-chip microfluidic potency assay is presented with improved functional predictive power and recapitulation of in vivo secretory responses compared to traditional approaches. Comparison of hMSC secretory responses to functional hMSC-medicated immune cell suppression demonstrates shortcomings of current surrogate potency markers and identifies on-chip microfluidic potency markers with improved functional predictive power compared to traditional planar methods. Furthermore, hMSC secretory performance achieved in the on-chip microfluidic system has improved similarity compared to an in vivo model. The results underscore the shortcomings of current culture practices and present a novel system with improved functional predictive power and hMSC physiological responses.
Keywords: biomaterials; cell therapies; mesenchymal stem/stromal cells; microfluidics; on-chip technologies.
© 2021 Wiley-VCH GmbH.
Figures
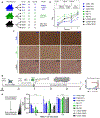

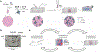



References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

