High rates of evolution preceded shifts to sex-biased gene expression in Leucadendron, the most sexually dimorphic angiosperms
- PMID: 34726596
- PMCID: PMC8635981
- DOI: 10.7554/eLife.67485
High rates of evolution preceded shifts to sex-biased gene expression in Leucadendron, the most sexually dimorphic angiosperms
Abstract
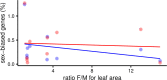
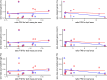
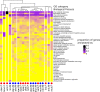
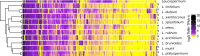
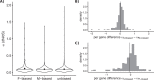
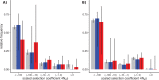
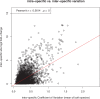
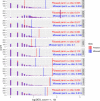
Differences between males and females are usually more subtle in dioecious plants than animals, but strong sexual dimorphism has evolved convergently in the South African Cape plant genus Leucadendron. Such sexual dimorphism in leaf size is expected largely to be due to differential gene expression between the sexes. We compared patterns of gene expression in leaves among 10 Leucadendron species across the genus. Surprisingly, we found no positive association between sexual dimorphism in morphology and the number or the percentage of sex-biased genes (SBGs). Sex bias in most SBGs evolved recently and was species specific. We compared rates of evolutionary change in expression for genes that were sex biased in one species but unbiased in others and found that SBGs evolved faster in expression than unbiased genes. This greater rate of expression evolution of SBGs, also documented in animals, might suggest the possible role of sexual selection in the evolution of gene expression. However, our comparative analysis clearly indicates that the more rapid rate of expression evolution of SBGs predated the origin of bias, and shifts towards bias were depleted in signatures of adaptation. Our results are thus more consistent with the view that sex bias is simply freer to evolve in genes less subject to constraints in expression level.
Keywords: Leucadendron; chromosomes; evolution; evolutionary biology; gene expression; plant; proteaceae; sex-biased gene expression; sexual dimorphism.
© 2021, Scharmann et al.
Conflict of interest statement
MS, AR, JP No competing interests declared
Figures



Similar articles
-
Positive selection and relaxed purifying selection contribute to rapid evolution of male-biased genes in a dioecious flowering plant.Elife. 2024 Feb 14;12:RP89941. doi: 10.7554/eLife.89941. Elife. 2024. PMID: 38353667 Free PMC article.
-
Positive selection and relaxed purifying selection contribute to rapid evolution of sex-biased genes in green seaweed Ulva.BMC Ecol Evol. 2025 May 9;25(1):44. doi: 10.1186/s12862-025-02382-y. BMC Ecol Evol. 2025. PMID: 40346481 Free PMC article.
-
Floral trait evolution associated with shifts between insect and wind pollination in the dioecious genus Leucadendron (Proteaceae).Evolution. 2016 Jan;70(1):126-39. doi: 10.1111/evo.12821. Epub 2015 Dec 9. Evolution. 2016. PMID: 26593965
-
Sexual dimorphism in flowering plants.J Exp Bot. 2013 Jan;64(1):67-82. doi: 10.1093/jxb/ers308. Epub 2012 Nov 25. J Exp Bot. 2013. PMID: 23183260 Review.
-
Genetic Architecture of Sexual Dimorphism in Humans.J Cell Physiol. 2015 Oct;230(10):2304-10. doi: 10.1002/jcp.24979. J Cell Physiol. 2015. PMID: 25740260 Review.
Cited by
-
Sex-Biased Gene Expression Under Sexually Antagonistic and Sex-Limited Selection.Mol Biol Evol. 2025 Jul 30;42(8):msaf178. doi: 10.1093/molbev/msaf178. Mol Biol Evol. 2025. PMID: 40729508 Free PMC article.
-
Novel Tri-Segmented Rhabdoviruses: A Data Mining Expedition Unveils the Cryptic Diversity of Cytorhabdoviruses.Viruses. 2023 Dec 10;15(12):2402. doi: 10.3390/v15122402. Viruses. 2023. PMID: 38140643 Free PMC article.
-
Positive selection and relaxed purifying selection contribute to rapid evolution of male-biased genes in a dioecious flowering plant.Elife. 2024 Feb 14;12:RP89941. doi: 10.7554/eLife.89941. Elife. 2024. PMID: 38353667 Free PMC article.
-
Selection drives convergent gene expression changes during transitions to co-sexuality in haploid sexual systems.Nat Ecol Evol. 2022 May;6(5):579-589. doi: 10.1038/s41559-022-01692-4. Epub 2022 Mar 21. Nat Ecol Evol. 2022. PMID: 35314785 Free PMC article.
-
Unveiling the genetic diversity of the genera Enamovirus and Polerovirus through data-driven virus discovery.Arch Virol. 2025 Mar 13;170(4):76. doi: 10.1007/s00705-025-06258-w. Arch Virol. 2025. PMID: 40080166
References
-
- Akaike H. In: Ternational Symposium on Information Theory. Petrov N, Csáki F, editors. Budapest, Hungary: Akadémia Kiadó; 1973. Information theory as an extension of the maximum likelihood principleB; pp. 267–281.
-
- Bachtrog D, Mank JE, Peichel CL, Kirkpatrick M, Otto SP, Ashman TL, Hahn MW, Kitano J, Mayrose I, Ming R, Perrin N, Ross L, Valenzuela N, Vamosi JC, Tree of Sex Consortium Sex determination: why so many ways of doing it? PLOS Biology. 2014;12:e1001899. doi: 10.1371/journal.pbio.1001899. - DOI - PMC - PubMed
Publication types
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources

