Deficiency of replication-independent DNA mismatch repair drives a 5-methylcytosine deamination mutational signature in cancer
- PMID: 34730999
- PMCID: PMC8565909
- DOI: 10.1126/sciadv.abg4398
Deficiency of replication-independent DNA mismatch repair drives a 5-methylcytosine deamination mutational signature in cancer
Abstract
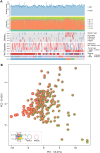
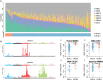
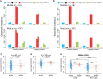
Multiple mutational signatures have been associated with DNA mismatch repair (MMR)–deficient cancers, but the mechanisms underlying their origin remain unclear. Here, using mutation data from cancer genomes, we identify a previously unknown function of MMR that is able to protect genomes from 5-methylcytosine (5mC) deamination–induced somatic mutations in a replication-independent manner. Cancers with deficiency of MMR proteins MSH2/MSH6 (MutSα) exhibit mutational signature contributions distinct from those that are deficient in MLH1/PMS2 (MutLα). This disparity arises from unrepaired 5mC deamination–induced mismatches rather than replicative DNA polymerase errors. In cancers with biallelic loss of MBD4 DNA glycosylase, repair of 5mC deamination damage is strongly associated with H3K36me3 chromatin, implicating MutSα as the essential factor in its repair. We thus uncover a noncanonical role of MMR in the protection against 5mC deamination–induced mutation in human cancers.
Figures



References
-
- Lynch H. T., Snyder C. L., Shaw T. G., Heinen C. D., Hitchins M. P., Milestones of Lynch syndrome: 1895-2015. Nat. Rev. Cancer 15, 181–194 (2015). - PubMed
-
- Hause R. J., Pritchard C. C., Shendure J., Salipante S. J., Classification and characterization of microsatellite instability across 18 cancer types. Nat. Med. 22, 1342–1350 (2016). - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

