Transcriptional and chromatin-based partitioning mechanisms uncouple protein scaling from cell size
- PMID: 34731644
- PMCID: PMC8642314
- DOI: 10.1016/j.molcel.2021.10.007
Transcriptional and chromatin-based partitioning mechanisms uncouple protein scaling from cell size
Abstract
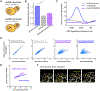
Biosynthesis scales with cell size such that protein concentrations generally remain constant as cells grow. As an exception, synthesis of the cell-cycle inhibitor Whi5 "sub-scales" with cell size so that its concentration is lower in larger cells to promote cell-cycle entry. Here, we find that transcriptional control uncouples Whi5 synthesis from cell size, and we identify histones as the major class of sub-scaling transcripts besides WHI5 by screening for similar genes. Histone synthesis is thereby matched to genome content rather than cell size. Such sub-scaling proteins are challenged by asymmetric cell division because proteins are typically partitioned in proportion to newborn cell volume. To avoid this fate, Whi5 uses chromatin-binding to partition similar protein amounts to each newborn cell regardless of cell size. Disrupting both Whi5 synthesis and chromatin-based partitioning weakens G1 size control. Thus, specific transcriptional and partitioning mechanisms determine protein sub-scaling to control cell size.
Keywords: cell cycle; cell size; cell size control; gene expression; scaling.
Copyright © 2021 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures




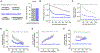
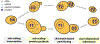
References
-
- Andrews BJ, and Herskowitz I (1989). Identification of a DNA binding factor involved in cell-cycle control of the yeast HO gene. Cell 57, 21–29. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

