QTL mapping in outbred tetraploid (and diploid) diallel populations
- PMID: 34740237
- PMCID: PMC8570786
- DOI: 10.1093/genetics/iyab124
QTL mapping in outbred tetraploid (and diploid) diallel populations
Abstract
Over the last decade, multiparental populations have become a mainstay of genetics research in diploid species. Our goal was to extend this paradigm to autotetraploids by developing software for quantitative trait locus (QTL) mapping in connected F1 populations derived from a set of shared parents. For QTL discovery, phenotypes are regressed on the dosage of parental haplotypes to estimate additive effects. Statistical properties of the model were explored by simulating half-diallel diploid and tetraploid populations with different population sizes and numbers of parents. Across scenarios, the number of progeny per parental haplotype (pph) largely determined the statistical power for QTL detection and accuracy of the estimated haplotype effects. Multiallelic QTL with heritability 0.2 were detected with 90% probability at 25 pph and genome-wide significance level 0.05, and the additive haplotype effects were estimated with over 90% accuracy. Following QTL discovery, the software enables a comparison of models with multiple QTL and nonadditive effects. To illustrate, we analyzed potato tuber shape in a half-diallel population with three tetraploid parents. A well-known QTL on chromosome 10 was detected, for which the inclusion of digenic dominance lowered the Deviance Information Criterion (DIC) by 17 points compared to the additive model. The final model also contained a minor QTL on chromosome 1, but higher-order dominance and epistatic effects were excluded based on the DIC. In terms of practical impacts, the software is already being used to select offspring based on the effect and dosage of particular haplotypes in breeding programs.
Keywords: MPP; Multiparental Populations; dominance; haplotypes; multiparental; polyploidy.
© The Author(s) 2021. Published by Oxford University Press on behalf of Genetics Society of America. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures
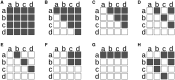
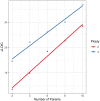
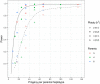
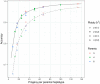

References
-
- Bink MCAM, Jansen J, Madduri M, Voorrips RE, Durel CE, et al.2014. Bayesian QTL analyses using pedigreed families of an outcrossing species, with application to fruit firmness in apple. Theor Appl Genet. 127:1073–1090. - PubMed
-
- Broman KW, Sen S.. 2009. A Guide to QTL Mapping with R/Qtl. Dordrecht, Holland: Springer.
Publication types
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources

