Predicting novel candidate human obesity genes and their site of action by systematic functional screening in Drosophila
- PMID: 34748544
- PMCID: PMC8575313
- DOI: 10.1371/journal.pbio.3001255
Predicting novel candidate human obesity genes and their site of action by systematic functional screening in Drosophila
Abstract
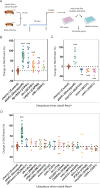
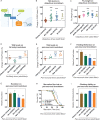
The discovery of human obesity-associated genes can reveal new mechanisms to target for weight loss therapy. Genetic studies of obese individuals and the analysis of rare genetic variants can identify novel obesity-associated genes. However, establishing a functional relationship between these candidate genes and adiposity remains a significant challenge. We uncovered a large number of rare homozygous gene variants by exome sequencing of severely obese children, including those from consanguineous families. By assessing the function of these genes in vivo in Drosophila, we identified 4 genes, not previously linked to human obesity, that regulate adiposity (itpr, dachsous, calpA, and sdk). Dachsous is a transmembrane protein upstream of the Hippo signalling pathway. We found that 3 further members of the Hippo pathway, fat, four-jointed, and hippo, also regulate adiposity and that they act in neurons, rather than in adipose tissue (fat body). Screening Hippo pathway genes in larger human cohorts revealed rare variants in TAOK2 associated with human obesity. Knockdown of Drosophila tao increased adiposity in vivo demonstrating the strength of our approach in predicting novel human obesity genes and signalling pathways and their site of action.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Comment in
-
Screening obesity genes in Drosophila.Lab Anim (NY). 2022 Jan;51(1):7. doi: 10.1038/s41684-021-00899-5. Lab Anim (NY). 2022. PMID: 34949849 No abstract available.
References
-
- Heymsfield SB, Wadden TA. Mechanisms, Pathophysiology, and Management of Obesity. N Engl J Med. 2017;376. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- MR/L003120/1/MRC_/Medical Research Council/United Kingdom
- WT102627/WT_/Wellcome Trust/United Kingdom
- 092096/WT_/Wellcome Trust/United Kingdom
- BTRU-2014-10024/DH_/Department of Health/United Kingdom
- RG/18/13/33946/BHF_/British Heart Foundation/United Kingdom
- 103792/WT_/Wellcome Trust/United Kingdom
- 206194/WT_/Wellcome Trust/United Kingdom
- WT210561/WT_/Wellcome Trust/United Kingdom
- C6946/A14492/CRUK_/Cancer Research UK/United Kingdom
- 207462/Z/17/Z/WT_/Wellcome Trust/United Kingdom
- M009017/MRC_/Medical Research Council/United Kingdom
- 208363/Z/17/Z/WT_/Wellcome Trust/United Kingdom
- RG/13/13/30194/BHF_/British Heart Foundation/United Kingdom
- SP/09/002/BHF_/British Heart Foundation/United Kingdom
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

