Infrared-spectroscopic, dynamic near-field microscopy of living cells and nanoparticles in water
- PMID: 34750511
- PMCID: PMC8576021
- DOI: 10.1038/s41598-021-01425-w
Infrared-spectroscopic, dynamic near-field microscopy of living cells and nanoparticles in water
Abstract
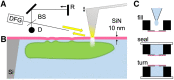
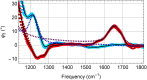
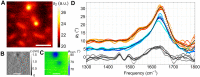
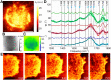
Infrared fingerprint spectra can reveal the chemical nature of materials down to 20-nm detail, far below the diffraction limit, when probed by scattering-type scanning near-field optical microscopy (s-SNOM). But this was impossible with living cells or aqueous processes as in corrosion, due to water-related absorption and tip contamination. Here, we demonstrate infrared s-SNOM of water-suspended objects by probing them through a 10-nm thick SiN membrane. This separator stretches freely over up to 250 µm, providing an upper, stable surface to the scanning tip, while its lower surface is in contact with the liquid and localises adhering objects. We present its proof-of-principle applicability in biology by observing simply drop-casted, living E. coli in nutrient medium, as well as living A549 cancer cells, as they divide, move and develop rich sub-cellular morphology and adhesion patterns, at 150 nm resolution. Their infrared spectra reveal the local abundances of water, proteins, and lipids within a depth of ca. 100 nm below the SiN membrane, as we verify by analysing well-defined, suspended polymer spheres and through model calculations. SiN-membrane based s-SNOM thus establishes a novel tool of live cell nano-imaging that returns structure, dynamics and chemical composition. This method should benefit the nanoscale analysis of any aqueous system, from physics to medicine.
© 2021. The Author(s).
Conflict of interest statement
F.K. was cofounder of Neaspec GmbH and has received compensation as a scientific advisor to attocube systems AG, companies producing scattering-type scanning near-field optical microscope systems such as the one used in this study. K.K. and T.G. are full-time and part-time employees, respectively, of attocube systems AG. E.B. declares no competing financial interest.
Figures



Similar articles
-
Nanoscale infrared absorption spectroscopy of individual nanoparticles enabled by scattering-type near-field microscopy.ACS Nano. 2011 Aug 23;5(8):6494-9. doi: 10.1021/nn2017638. Epub 2011 Jul 27. ACS Nano. 2011. PMID: 21770439
-
Nanoscale Optical Microscopy and Spectroscopy Using Near-Field Probes.Annu Rev Chem Biomol Eng. 2018 Jun 7;9:365-387. doi: 10.1146/annurev-chembioeng-060817-084150. Epub 2018 Mar 29. Annu Rev Chem Biomol Eng. 2018. PMID: 29596000 Review.
-
Graphene-Based Platform for Infrared Near-Field Nanospectroscopy of Water and Biological Materials in an Aqueous Environment.ACS Nano. 2015 Aug 25;9(8):7968-75. doi: 10.1021/acsnano.5b01184. Epub 2015 Aug 5. ACS Nano. 2015. PMID: 26223158
-
Infrared spectroscopic mapping of single nanoparticles and viruses at nanoscale resolution.Nano Lett. 2006 Jul;6(7):1307-10. doi: 10.1021/nl0610836. Nano Lett. 2006. PMID: 16834401
-
Infrared Imaging and Spectroscopy Beyond the Diffraction Limit.Annu Rev Anal Chem (Palo Alto Calif). 2015;8:101-26. doi: 10.1146/annurev-anchem-071114-040435. Epub 2015 May 18. Annu Rev Anal Chem (Palo Alto Calif). 2015. PMID: 26001952 Review.
Cited by
-
Nanoscale Mechanical Manipulation of Ultrathin SiN Membranes Enabling Infrared Near-Field Microscopy of Liquid-Immersed samples.Small. 2024 Nov;20(47):e2402568. doi: 10.1002/smll.202402568. Epub 2024 Aug 15. Small. 2024. PMID: 39148207 Free PMC article.
-
Synchrotron nano-FTIR spectroscopy for probing anticancer drugs at subcellular scale.Sci Rep. 2024 Jul 26;14(1):17166. doi: 10.1038/s41598-024-67386-y. Sci Rep. 2024. PMID: 39060284 Free PMC article.
-
Transient infrared nanoscopy resolves the millisecond photoswitching dynamics of single lipid vesicles in water.Nat Commun. 2025 Jul 1;16(1):6033. doi: 10.1038/s41467-025-61341-9. Nat Commun. 2025. PMID: 40593745 Free PMC article.
-
High-fidelity nano-FTIR spectroscopy by on-pixel normalization of signal harmonics.Nanophotonics. 2021 Dec 13;11(2):377-390. doi: 10.1515/nanoph-2021-0565. eCollection 2022 Jan. Nanophotonics. 2021. PMID: 39633877 Free PMC article.
-
Scanning Probe Nano-Infrared Imaging and Spectroscopy of Biochemical and Natural Materials.Small Sci. 2024 Sep 26;4(11):2400297. doi: 10.1002/smsc.202400297. eCollection 2024 Nov. Small Sci. 2024. PMID: 40213449 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

