Conventional rigid 2D substrates cause complex contractile signals in monolayers of human induced pluripotent stem cell-derived cardiomyocytes
- PMID: 34761809
- PMCID: PMC9299844
- DOI: 10.1113/JP282228
Conventional rigid 2D substrates cause complex contractile signals in monolayers of human induced pluripotent stem cell-derived cardiomyocytes
Abstract
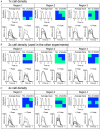
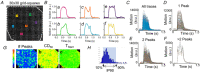
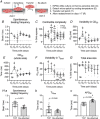
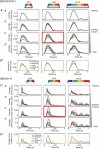
Human induced pluripotent stem cell-derived cardiomyocytes (hiPSC-CM) in monolayers interact mechanically via cell-cell and cell-substrate adhesion. Spatiotemporal features of contraction were analysed in hiPSC-CM monolayers (1) attached to glass or plastic (Young's modulus (E) >1 GPa), (2) detached (substrate-free) and (3) attached to a flexible collagen hydrogel (E = 22 kPa). The effects of isoprenaline on contraction were compared between rigid and flexible substrates. To clarify the underlying mechanisms, further gene expression and computational studies were performed. HiPSC-CM monolayers exhibited multiphasic contractile profiles on rigid surfaces in contrast to hydrogels, substrate-free cultures or single cells where only simple twitch-like time-courses were observed. Isoprenaline did not change the contraction profile on either surface, but its lusitropic and chronotropic effects were greater in hydrogel compared with glass. There was no significant difference between stiff and flexible substrates in regard to expression of the stress-activated genes NPPA and NPPB. A computational model of cell clusters demonstrated similar complex contractile interactions on stiff substrates as a consequence of cell-to-cell functional heterogeneity. Rigid biomaterial surfaces give rise to unphysiological, multiphasic contractions in hiPSC-CM monolayers. Flexible substrates are necessary for normal twitch-like contractility kinetics and interpretation of inotropic interventions. KEY POINTS: Spatiotemporal contractility analysis of human induced pluripotent stem cell-derived cardiomyocyte (hiPSC-CM) monolayers seeded on conventional, rigid surfaces (glass or plastic) revealed the presence of multiphasic contraction patterns across the monolayer with a high variability, despite action potentials recorded in the same areas being identical. These multiphasic patterns are not present in single cells, in detached monolayers or in monolayers seeded on soft substrates such as a hydrogel, where only 'twitch'-like transients are observed. HiPSC-CM monolayers that display a high percentage of regions with multiphasic contraction have significantly increased contractile duration and a decreased lusotropic drug response. There is no indication that the multiphasic contraction patterns are associated with significant activation of the stress-activated NPPA or NPPB signalling pathways. A computational model of cell clusters supports the biological findings that the rigid surface and the differential cell-substrate adhesion underly multiphasic contractile behaviour of hiPSC-CMs.
Keywords: cardiac physiology; mathematical model; pharmaceutical assay; recombinant collagen polymer; substrate rigidity.
© 2021 The Authors. The Journal of Physiology published by John Wiley & Sons Ltd on behalf of The Physiological Society.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures








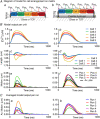
References
-
- Bers DM (2002). Cardiac excitation‐contraction coupling. Nature 415, 198–205. - PubMed
-
- Blinova K, Dang Q, Millard D, Smith G, Pierson J, Guo L, Brock M, Lu HR, Kraushaar U, Zeng H, Shi H, Zhang X, Sawada K, Osada T, Kanda Y, Sekino Y, Pang L, Feaster TK, Kettenhofen R, Stockbridge N, Strauss DG & Gintant G (2018). International multisite study of human‐induced pluripotent stem cell‐derived cardiomyocytes for drug proarrhythmic potential assessment. Cell Rep 24, 3582–3592. - PMC - PubMed
-
- Cerignoli F, Charlot D, Whittaker R, Ingermanson R, Gehalot P, Savchenko A, Gallacher DJ, Towart R, Price JH, McDonough PM & Mercola M (2012). High throughput measurement of Ca2⁺ dynamics for drug risk assessment in human stem cell‐derived cardiomyocytes by kinetic image cytometry. J Pharmacol Toxicol Methods 66, 246–256. - PMC - PubMed
-
- Chen G, Li S, Karakikes I, Ren L, Chow MZ, Chopra A, Keung W, Yan B, Chan CW, Costa KD, Kong CW, Hajjar RJ, Chen CS & Li RA (2015). Phospholamban as a crucial determinant of the inotropic response of human pluripotent stem cell‐derived ventricular cardiomyocytes and engineered 3‐dimensional tissue constructs. Circ Arrhythm Electrophysiol 8, 193–202. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

