Culture-Independent Exploration of the Hypersaline Ecosystem Indicates the Environment-Specific Microbiome Evolution
- PMID: 34777269
- PMCID: PMC8581802
- DOI: 10.3389/fmicb.2021.686549
Culture-Independent Exploration of the Hypersaline Ecosystem Indicates the Environment-Specific Microbiome Evolution
Abstract
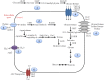
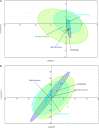
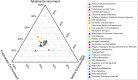
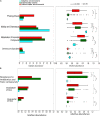
Sambhar Salt Lake, situated in the state of Rajasthan, India is a unique temperate hypersaline ecosystem. Exploration of the salt lake microbiome will enable us to understand microbiome functioning in nutrient-deprived extreme conditions, as well as enrich our understanding of the environment-specific microbiome evolution. The current study has been designed to explore the Sambhar Salt Lake microbiome with a culture-independent multi-omics approach to define its metagenomic features and prevalent metabolic functionaries. The rRNA feature and protein feature-based phylogenetic reconstruction synchronously (R = 0.908) indicated the dominance of the archaea (Euryarchaeota) and bacteria (Firmicutes, Proteobacteria, Bacteroidetes, and Actinobacteria). Metabolic reconstruction identified selective enrichment of the protein features associated with energy harvesting and stress tolerance (osmotic, oxidative, metal/metalloid, heat/cold, antibiotic, and desiccation). Metabolites identified with metabolome analysis confirmed physiological adaptation of the lake microbiome within a hypersaline and nutrient-deprived environment. Comparative metagenomics of the 212 metagenomes representing freshwater, alkaline, and saline ecosystem microbiome indicated the selective enrichment of the microbial groups and genetic features. The current study elucidates microbiome functioning within the nutrient-deprived harsh ecosystems. In summary, the current study harnessing the strength of multi-omics and comparative metagenomics indicates the environment-specific microbiome evolution.
Keywords: extremophiles; hypersaline lake; metagenome; microbiome; microbiome physiology; multi-omics analysis; stress response and adaptation.
Copyright © 2021 Mehta, Yadav, Ahmed, Goyal, Pandey and Chauhan.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
-
- Achberger A. M., Michaud A. B., Vick-Majors T. J., Christner B. C., Skidmore M. L., Priscu J. C., et al. (2017). “Microbiology of subglacial environments,” in Psychrophiles: From Biodiversity to Biotechnology, ed. Margesin R. (Cham: Springer; ), 83–110.
-
- Arun N., Singh D. P. (2014). Chromium (VI) induced oxidative stress in halotolerant alga Dunaliellasalina and D. tertiolecta isolated from Sambhar salt lake of Rajasthan (India). Cell. Mol. Biol. 60 90–96. - PubMed
LinkOut - more resources
Full Text Sources

