CPVL promotes glioma progression via STAT1 pathway inhibition through interactions with the BTK/p300 axis
- PMID: 34784299
- PMCID: PMC8783677
- DOI: 10.1172/jci.insight.146362
CPVL promotes glioma progression via STAT1 pathway inhibition through interactions with the BTK/p300 axis
Abstract
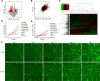
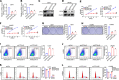
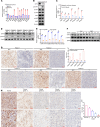
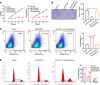
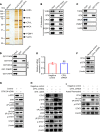
CPVL (carboxypeptidase, vitellogenic-like) is a serine carboxypeptidase that was first characterized in human macrophages. However, the function of CPVL remains unclear in a variety of tumors. The quantitative PCR (qPCR), Western blotting, and IHC assays were utilized to measure the CPVL expression. CPVL was significantly upregulated in glioma cells and tissues compared with normal cells and tissues, respectively. Moreover, high CPVL expression was correlated with advanced clinical grade and poor prognosis. Silencing of CPVL promoted glioma cell apoptosis, and it inhibited cell proliferation and tumorigenicity in vitro and in vivo. Ingenuity Pathway Analysis (IPA) demonstrated that CPVL silencing activated the IFN-γ/STAT1 signaling pathway, thereby inducing glioma cell apoptosis. Mechanistically, immunopurification, mass spectrometry, IP, and glutathione S-transferase (GST) pull-down experiments elucidated that CPVL physically interacts with Bruton's tyrosine kinase (BTK) and downregulates the STAT1 phosphorylation through promoting p300-mediated STAT1 acetylation. Our findings reveal the crucial role of CPVL in promoting the progression of glioma through suppressing STAT1 phosphorylation. CPVL might serve as a potential prognostic biomarker and therapeutic target for the treatment of glioma.
Keywords: Oncogenes; Oncology.
Figures


Similar articles
-
Carboxypeptidase Vitellogenic-Like Promotes the Proliferation and Metastasis of Osteosarcoma through the TGF-β/Smad Signaling Pathway.Ann Clin Lab Sci. 2023 Nov;53(6):890-904. Ann Clin Lab Sci. 2023. PMID: 38182149
-
Activation of STAT1 by the FRK tyrosine kinase is associated with human glioma growth.J Neurooncol. 2019 May;143(1):35-47. doi: 10.1007/s11060-019-03143-w. Epub 2019 Apr 16. J Neurooncol. 2019. PMID: 30993511
-
Cloning and characterization of CPVL, a novel serine carboxypeptidase, from human macrophages.Genomics. 2001 Mar 15;72(3):243-51. doi: 10.1006/geno.2000.6484. Genomics. 2001. PMID: 11401439
-
Carboxypeptidase vitellogenic like facilitates resistance to CDK4/6 inhibitors in breast cancer.Thorac Cancer. 2023 Apr;14(11):983-991. doi: 10.1111/1759-7714.14829. Epub 2023 Feb 24. Thorac Cancer. 2023. PMID: 36825764 Free PMC article.
-
The IFNgamma-induced STAT1-CBP/P300 association, required for a normal response to the cytokine, is disrupted in Brucella-infected macrophages.Microb Pathog. 2009 Feb;46(2):88-97. doi: 10.1016/j.micpath.2008.10.011. Epub 2008 Nov 13. Microb Pathog. 2009. PMID: 19041714
Cited by
-
Dissecting gastric cancer heterogeneity and exploring therapeutic strategies using bulk and single-cell transcriptomic analysis and experimental validation of tumor microenvironment and metabolic interplay.Front Pharmacol. 2024 Jun 19;15:1355269. doi: 10.3389/fphar.2024.1355269. eCollection 2024. Front Pharmacol. 2024. PMID: 38962317 Free PMC article.
-
BTK Expression Level Prediction and the High-Grade Glioma Prognosis Using Radiomic Machine Learning Models.J Imaging Inform Med. 2024 Aug;37(4):1359-1374. doi: 10.1007/s10278-024-01026-9. Epub 2024 Feb 21. J Imaging Inform Med. 2024. PMID: 38381384 Free PMC article.
-
OTUB1 Promotes Glioblastoma Growth by Inhibiting the JAK2/STAT1 Signaling Pathway.J Cancer. 2024 Jun 24;15(14):4566-4576. doi: 10.7150/jca.96360. eCollection 2024. J Cancer. 2024. PMID: 39006090 Free PMC article.
-
Complementary CRISPR genome-wide genetic screens in PARP10-knockout and overexpressing cells identify synthetic interactions for PARP10-mediated cellular survival.Oncotarget. 2022 Sep 28;13:1078-1091. doi: 10.18632/oncotarget.28277. eCollection 2022. Oncotarget. 2022. PMID: 36187556 Free PMC article.
-
Single-cell Atlas reveals core function of CPVL/MSR1 expressing macrophages in the prognosis of triple-negative breast cancer.Front Immunol. 2024 Dec 24;15:1501009. doi: 10.3389/fimmu.2024.1501009. eCollection 2024. Front Immunol. 2024. PMID: 39776914 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

