The bHLH transcription factor AhbHLH112 improves the drought tolerance of peanut
- PMID: 34784902
- PMCID: PMC8594184
- DOI: 10.1186/s12870-021-03318-6
The bHLH transcription factor AhbHLH112 improves the drought tolerance of peanut
Abstract
Background: Basic helix-loop-helix (bHLH) transcription factors (TFs) are one of the largest gene families in plants. They regulate gene expression through interactions with specific motifs in target genes. bHLH TFs are not only universally involved in plant growth but also play an important role in plant responses to abiotic stress. However, most members of this family have not been functionally characterized.
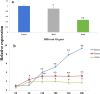
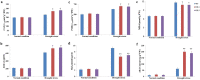
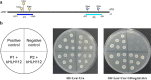
Results: Here, we characterized the function of a bHLH TF in the peanut, AhHLH112, in response to drought stress. AhHLH112 is localized in the nucleus and it was induced by drought stress. The overexpression of this gene improves the drought tolerance of transgenic plants both in seedling and adult stages. Compared to wild-type plants, the transgenic plants accumulated less reactive oxygen species (ROS), accompanied by increased activity and transcript levels of antioxidant enzymes (superoxide dismutase, peroxidase and catalase). In addition, the WT plants demonstrated higher MDA concentration levels and higher water loss rate than the transgenic plants under drought treatment. The Yeast one-hybrid result also demonstrates that AhbHLH112 directly and specifically binds to and activates the promoter of the peroxidase (POD) gene. Besides, overexpression of AhHLH112 improved ABA level under drought condition, and elevated the expression of genes associated with ABA biosynthesis and ABA responding, including AtNCED3 and AtRD29A.
Conclusions: Drawing on the results of our experiments, we propose that, by improving ROS-scavenging ability, at least in part through the regulation of POD -mediated H2O2 homeostasis, and possibly participates in ABA-dependent stress-responding pathway, AhbHLH112 acts as a positive factor in drought stress tolerance.
Keywords: Basic helix–loop–helix transcription factors; Drought stress; Peanut; ROS homeostasis; Transcriptional regulation.
© 2021. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




References
-
- Zhao X, Li C, Wan S, Zhang T, Yan C, Shan S. Transcriptomic analysis and discovery of genes in the response of Arachis hypogaea to drought stress. Mol Biol Rep. 2018;45:119–131. - PubMed
-
- Bertioli DJ, Cannon SB, Froenicke L, Huang G, Farmer AD, Cannon EKS, et al. The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat Genet. 2016;48:438–446. - PubMed
-
- Bertioli DJ, Jenkins J, Clevenger J, Dudchenko O, Gao D, Seijo G, et al. The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nat Genet. 2019;51:877–884. - PubMed
-
- DaMatta FM. Exploring drought tolerance in coffee: a physiologicalapproach with some insights for plant breeding. Braz J Plant Physiol. 2004;16:1–6.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

