Disparity-filtered differential correlation network analysis: a case study on CRC metabolomics
- PMID: 34792303
- PMCID: PMC8709737
- DOI: 10.1515/jib-2021-0030
Disparity-filtered differential correlation network analysis: a case study on CRC metabolomics
Abstract
Differential network analysis has become a widely used technique to investigate changes of interactions among different conditions. Although the relationship between observed interactions and biochemical mechanisms is hard to establish, differential network analysis can provide useful insights about dysregulated pathways and candidate biomarkers. The available methods to detect differential interactions are heterogeneous and often rely on assumptions that are unrealistic in many applications. To address these issues, we develop a novel method for differential network analysis, using the so-called disparity filter as network reduction technique. In addition, we propose a classification model based on the inferred network interactions. The main novelty of this work lies in its ability to preserve connections that are statistically significant with respect to a null model without favouring any resolution scale, as a hard threshold would do, and without Gaussian assumptions. The method was tested using a published metabolomic dataset on colorectal cancer (CRC). Detected hub metabolites were consistent with recent literature and the classifier was able to distinguish CRC from polyp and healthy subjects with great accuracy. In conclusion, the proposed method provides a new simple and effective framework for the identification of differential interaction patterns and improves the biological interpretation of metabolomics data.
Keywords: colorectal cancer (CRC); differential network analysis; disparity filter; metabolomics.
© 2021 Silvia Sabatini et al., published by De Gruyter, Berlin/Boston.
Figures
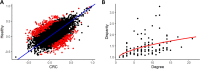

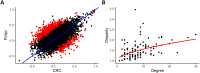

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
