Epigenetically mismatched parental centromeres trigger genome elimination in hybrids
- PMID: 34797718
- PMCID: PMC8604413
- DOI: 10.1126/sciadv.abk1151
Epigenetically mismatched parental centromeres trigger genome elimination in hybrids
Abstract
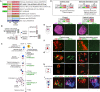
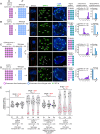
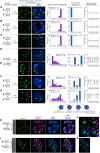
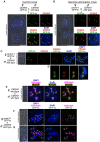
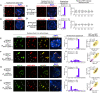
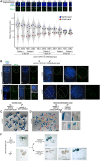
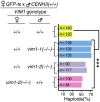
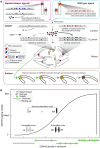
Wide crosses result in postzygotic elimination of one parental chromosome set, but the mechanisms that result in such differential fate are poorly understood. Here, we show that alterations of centromeric histone H3 (CENH3) lead to its selective removal from centromeres of mature Arabidopsis eggs and early zygotes, while wild-type CENH3 persists. In the hybrid zygotes and embryos, CENH3 and essential centromere proteins load preferentially on the CENH3-rich centromeres of the wild-type parent, while CENH3-depleted centromeres fail to reconstitute new CENH3-chromatin and the kinetochore and are frequently lost. Genome elimination is opposed by E3 ubiquitin ligase VIM1. We propose a model based on cooperative binding of CENH3 to chromatin to explain the differential CENH3 loading rates. Thus, parental CENH3 polymorphisms result in epigenetically distinct centromeres that instantiate a strong mating barrier and produce haploids.
Figures


Similar articles
-
Naturally occurring differences in CENH3 affect chromosome segregation in zygotic mitosis of hybrids.PLoS Genet. 2015 Jan 26;11(1):e1004970. doi: 10.1371/journal.pgen.1004970. eCollection 2015 Jan. PLoS Genet. 2015. PMID: 25622028 Free PMC article.
-
Plasticity of parental CENH3 incorporation into the centromeres in wheat × barley F1 hybrids.Front Plant Sci. 2024 Jan 19;15:1324817. doi: 10.3389/fpls.2024.1324817. eCollection 2024. Front Plant Sci. 2024. PMID: 38313805 Free PMC article.
-
Loss of centromeric histone H3 (CENH3) from centromeres precedes uniparental chromosome elimination in interspecific barley hybrids.Proc Natl Acad Sci U S A. 2011 Aug 16;108(33):E498-505. doi: 10.1073/pnas.1103190108. Epub 2011 Jul 11. Proc Natl Acad Sci U S A. 2011. PMID: 21746892 Free PMC article.
-
Centromere Size and Its Relationship to Haploid Formation in Plants.Mol Plant. 2018 Mar 5;11(3):398-406. doi: 10.1016/j.molp.2017.12.009. Epub 2017 Dec 19. Mol Plant. 2018. PMID: 29277426 Review.
-
Centromere histone H3- and phospholipase-mediated haploid induction in plants.Plant Methods. 2019 Apr 26;15:42. doi: 10.1186/s13007-019-0429-5. eCollection 2019. Plant Methods. 2019. PMID: 31057661 Free PMC article. Review.
Cited by
-
A centromere's obsession with transposons.Nat Plants. 2024 Sep;10(9):1282-1283. doi: 10.1038/s41477-024-01724-w. Nat Plants. 2024. PMID: 39223306 No abstract available.
-
Centromeres drive and take a break.Chromosome Res. 2025 Aug 4;33(1):17. doi: 10.1007/s10577-025-09777-z. Chromosome Res. 2025. PMID: 40759764 Free PMC article. Review.
-
Driving lessons: a brief (personal) history of centromere drive.Genetics. 2022 Nov 30;222(4):iyac155. doi: 10.1093/genetics/iyac155. Genetics. 2022. PMID: 39255401 Free PMC article. No abstract available.
-
Heat stress impairs centromere structure and segregation of meiotic chromosomes in Arabidopsis.Elife. 2024 Apr 17;12:RP90253. doi: 10.7554/eLife.90253. Elife. 2024. PMID: 38629825 Free PMC article.
-
Synthetic maize centromeres transmit chromosomes across generations.Nat Plants. 2023 Mar;9(3):433-441. doi: 10.1038/s41477-023-01370-8. Epub 2023 Mar 16. Nat Plants. 2023. PMID: 36928774
References
-
- Ishii T., Karimi-Ashtiyani R., Houben A., Haploidization via chromosome elimination: Means and mechanisms. Annu. Rev. Plant Biol. 67, 421–438 (2016). - PubMed
-
- Ravi M., Chan S. W. L., Haploid plants produced by centromere-mediated genome elimination. Nature 464, 615–618 (2010). - PubMed
-
- Lv J., Yu K., Wei J., Gui H., Liu C., Liang D., Wang Y., Zhou H., Carlin R., Rich R., Lu T., Que Q., Wang W. C., Zhang X., Kelliher T., Generation of paternal haploids in wheat by genome editing of the centromeric histone CENH3. Nat. Biotechnol. 38, 1397–1401 (2020). - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

