Co-targeting BET bromodomain BRD4 and RAC1 suppresses growth, stemness and tumorigenesis by disrupting the c-MYC-G9a-FTH1axis and downregulating HDAC1 in molecular subtypes of breast cancer
- PMID: 34803511
- PMCID: PMC8579449
- DOI: 10.7150/ijbs.62236
Co-targeting BET bromodomain BRD4 and RAC1 suppresses growth, stemness and tumorigenesis by disrupting the c-MYC-G9a-FTH1axis and downregulating HDAC1 in molecular subtypes of breast cancer
Abstract
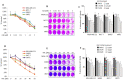
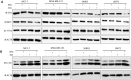
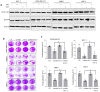
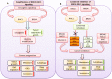
BET bromodomain BRD4 and RAC1 oncogenes are considered important therapeutic targets for cancer and play key roles in tumorigenesis, survival and metastasis. However, combined inhibition of BRD4-RAC1 signaling pathways in different molecular subtypes of breast cancer including luminal-A, HER-2 positive and triple-negative breast (TNBC) largely remains unknown. Here, we demonstrated a new co-targeting strategy by combined inhibition of BRD4-RAC1 oncogenic signaling in different molecular subtypes of breast cancer in a context-dependent manner. We show that combined treatment of JQ1 (inhibitor of BRD4) and NSC23766 (inhibitor of RAC1) suppresses cell growth, clonogenic potential, cell migration and mammary stem cells expansion and induces autophagy and cellular senescence in molecular subtypes of breast cancer cells. Mechanistically, JQ1/NSC23766 combined treatment disrupts MYC/G9a axis and subsequently enhances FTH1 to exert antitumor effects. Furthermore, combined treatment targets HDAC1/Ac-H3K9 axis, thus suggesting a role of this combination in histone modification and chromatin modeling. C-MYC depletion and co-treatment with vitamin-C sensitizes different molecular subtypes of breast cancer cells to JQ1/NSC23766 combination and further reduces cell growth, cell migration and mammosphere formation. Importantly, co-targeting RAC1-BRD4 suppresses breast tumor growth in vivo using xenograft mouse model. Clinically, RAC1 and BRD4 expression positively correlates in breast cancer patient's samples and show high expression patterns across different molecular subtypes of breast cancer. Both RAC1 and BRD4 proteins predict poor survival in breast cancer patients. Taken together, our results suggest that combined inhibition of BRD4-RAC1 pathways represents a novel and potential therapeutic approach in different molecular subtypes of breast cancer and highlights the importance of co-targeting RAC1-BRD4 signaling in breast tumorigenesis via disruption of C-MYC/G9a/FTH1 axis and down regulation of HDAC1.
Keywords: BET bromodomain; BRD4; Breast cancer; FTH1; G9a; HDAC1; JQ1; NSC23766; RAC1; c-MYC.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures







References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

