The Complete Plastid Genomes of Seven Sargassaceae Species and Their Phylogenetic Analysis
- PMID: 34804089
- PMCID: PMC8602799
- DOI: 10.3389/fpls.2021.747036
The Complete Plastid Genomes of Seven Sargassaceae Species and Their Phylogenetic Analysis
Abstract
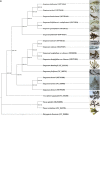
Sargassum is one of the most important genera of the family Sargassaceae in brown algae and is used to produce carrageenan, mannitol, iodine, and other economic substances. Here, seven complete plastid genomes of Sargassum ilicifolium var. conduplicatum, S. graminifolium, S. phyllocystum, S. muticum, S. feldmannii, S. mcclurei, and S. henslowianum were assembled using next-generation sequencing. The sizes of the seven circular genomes ranged from 124,258 to 124,563 bp, with two inverted regions and the same set of plastid genes, including 139 protein-coding genes (PCGs), 28 transfer (t)RNAs, and 6 ribosomal (r)RNAs. Compared with the other five available plastid genomes of Fucales, 136 PCGs were conserved, with two common ones shared with Coccophora langsdorfii, and one with S. fusiforme and S. horneri. The co-linear analysis identified two inversions of trnC(gca) and trnN(gtt) in ten Sargassum species, against S. horneri and C. langsdorfii. The phylogenetic analysis based on the plastid genomes of 55 brown algae (Phaeophyceae) showed four clades, whose ancient ancestor lived around 201.42 million years ago (Mya), and the internal evolutionary branches in Fucales started to be formed 92.52 Mya, while Sargassum species were divided into two subclades 14.33 Mya. Our novel plastid genomes provided evidence for the speciation of brown algae and plastid genomic evolution events.
Keywords: Sargassaceae; brown algae; co-linear analysis; comparative analysis; phylogenetic analysis; plastid genome.
Copyright © 2021 Li, Jia, Zhang, Jia, Liu, Qu and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Complete mitochondrial genome of the invasive brown alga Sargassum muticum (Sargassaceae, Phaeophyceae).Mitochondrial DNA A DNA Mapp Seq Anal. 2016;27(2):1129-30. doi: 10.3109/19401736.2014.933333. Epub 2015 Sep 4. Mitochondrial DNA A DNA Mapp Seq Anal. 2016. PMID: 24983154
-
Plastid and mitochondrial genomes of Coccophora langsdorfii (Fucales, Phaeophyceae) and the utility of molecular markers.PLoS One. 2017 Nov 2;12(11):e0187104. doi: 10.1371/journal.pone.0187104. eCollection 2017. PLoS One. 2017. PMID: 29095864 Free PMC article.
-
Complete plastid genome of an ecologically important brown alga Sargassum thunbergii (Fucales, Phaeophyceae).Mar Genomics. 2016 Aug;28:17-20. doi: 10.1016/j.margen.2016.03.003. Epub 2016 Mar 21. Mar Genomics. 2016. PMID: 27012360
-
Complete mitochondrial genome of the brown alga Sargassum fusiforme (Sargassaceae, Phaeophyceae): genome architecture and taxonomic consideration.Mitochondrial DNA A DNA Mapp Seq Anal. 2016;27(2):1158-60. doi: 10.3109/19401736.2014.936417. Epub 2015 Sep 4. Mitochondrial DNA A DNA Mapp Seq Anal. 2016. PMID: 24989050
-
Diversified Chemical Structures and Bioactivities of the Chemical Constituents Found in the Brown Algae Family Sargassaceae.Mar Drugs. 2024 Jan 24;22(2):59. doi: 10.3390/md22020059. Mar Drugs. 2024. PMID: 38393030 Free PMC article. Review.
Cited by
-
Organellar Genomes of Sargassum hemiphyllum var. chinense Provide Insight into the Characteristics of Phaeophyceae.Int J Mol Sci. 2024 Aug 6;25(16):8584. doi: 10.3390/ijms25168584. Int J Mol Sci. 2024. PMID: 39201271 Free PMC article.
References
-
- Aaron-Amper J., Largo D. B., Handugan R. B., Nini J. L., Gulayan S. J. (2020). Culture of the tropical brown seaweed Sargassum aquifolium: from hatchery to field out-planting. Aquac. Rep. 16:100265. 10.1016/j.aqrep.2019.100265 - DOI
-
- Bermúdez Y. G., Rico I. L. R., Bermúdez O. G., Guibal E. (2011). Nickel biosorption using Gracilaria caudata and Sargassum muticum. Chem. Eng. J. 166 122–131. 10.1016/j.cej.2010.10.038 - DOI
-
- Bock R., Knoop V. (2012). Genomics of Chloroplasts and Mitochondria. Netherlands: Springer.
-
- Bringloe T. T., Starko S., Wade R. M., Vieira C., Kawai H., Clerck O. D., et al. (2020). Phylogeny and Evolution of the Brown Algae. Crit. Rev. Plant Sci. 39 281–321. 10.1080/07352689.2020.1787679 - DOI
-
- Cao M. (2015). Analysis Of Genome Structure Features Of Pyropia Haitanensis And Functional Genomics of Bangia fuscopurpurea. Qingdao: Ocean University of China.
LinkOut - more resources
Full Text Sources

