Diverse mechanisms of centromere specification
- PMID: 34813757
- PMCID: PMC8820161
- DOI: 10.1016/j.cub.2021.09.083
Diverse mechanisms of centromere specification
Abstract
The centromere performs a universally conserved function, to accurately partition genetic information upon cell division. Yet, centromeres are among the most rapidly evolving regions of the genome and are bound by a varying assortment of centromere-binding factors that are themselves highly divergent at the protein-sequence level. A common thread in most species is the dependence on the centromere-specific histone variant CENP-A for the specification of the centromere site. However, CENP-A is not universally required in all species or cell types, making the identification of a general mechanism for centromere specification challenging. In this review, we examine our current understanding of the mechanisms of centromere specification in CENP-A-dependent and independent systems, focusing primarily on recent work.
Copyright © 2021 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures

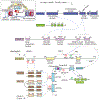
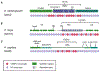
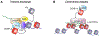
References
-
- Santaguida S, and Amon A (2015). Short- and long-term effects of chromosome mis-segregation and aneuploidy. Nature Reviews Molecular Cell Biology 16, 473–485. - PubMed
-
- Murillo-Pineda M, and Jansen LET (2020). Genetics, epigenetics and back again: Lessons learned from neocentromeres. Experimental Cell Research 389, 111909. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

