Urinary peptidomics reveals proteases involved in idiopathic membranous nephropathy
- PMID: 34819020
- PMCID: PMC8613922
- DOI: 10.1186/s12864-021-08155-3
Urinary peptidomics reveals proteases involved in idiopathic membranous nephropathy
Abstract
Background: Idiopathic membranous nephropathy (IMN) is a cause of nephrotic syndrome that is increasing in incidence but has unclear pathogenesis. Urinary peptidomics is a promising technology for elucidating molecular mechanisms underlying diseases. Dysregulation of the proteolytic system is implicated in various diseases. Here, we aimed to conduct urinary peptidomics to identify IMN-related proteases.
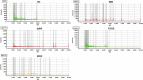
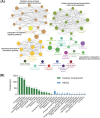
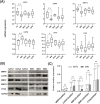
Results: Peptide fingerprints indicated differences in naturally produced urinary peptide components among 20 healthy individuals, 22 patients with IMN, and 15 patients with other kidney diseases. In total, 1,080 peptide-matched proteins were identified, 279 proteins differentially expressed in the urine of IMN patients were screened, and 32 proteases were predicted; 55 of the matched proteins were also differentially expressed in the kidney tissues of IMN patients, and these were mainly involved in the regulation of proteasome-, lysosome-, and actin cytoskeleton-related signaling pathways. The 32 predicted proteases showed abnormal expression in the glomeruli of IMN patients based on Gene Expression Omnibus databases. Western blot revealed abnormal expression of calpain, matrix metalloproteinase 14, and cathepsin S in kidney tissues of patients with IMN.
Conclusions: This work shown the calpain/matrix metalloproteinase/cathepsin axis might be dysregulated in IMN. Our study is the first to systematically explore the role of proteases in IMN by urinary peptidomics, which are expected to facilitate discovery of better biomarkers for IMN.
Keywords: Idiopathic membranous nephropathy; Proteases; Urinary peptidomics.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Differential Expression of Urinary Exosomal Small RNAs in Idiopathic Membranous Nephropathy.Biomed Res Int. 2020 Dec 29;2020:3170927. doi: 10.1155/2020/3170927. eCollection 2020. Biomed Res Int. 2020. PMID: 33457405 Free PMC article.
-
Urinary exosomal hsa_circ_0001250 as a novel diagnostic biomarker of idiopathic membranous nephropathy.J Transl Med. 2022 Dec 19;20(1):607. doi: 10.1186/s12967-022-03784-y. J Transl Med. 2022. PMID: 36536378 Free PMC article.
-
Application of miR-193a/WT1/PODXL axis to estimate risk and prognosis of idiopathic membranous nephropathy.Ren Fail. 2019 Nov;41(1):704-717. doi: 10.1080/0886022X.2019.1642210. Ren Fail. 2019. PMID: 31352863 Free PMC article.
-
Advances in antigens associated with Idiopathic Membranous Nephropathy.J Formos Med Assoc. 2021 Nov;120(11):1941-1948. doi: 10.1016/j.jfma.2021.06.014. Epub 2021 Jul 7. J Formos Med Assoc. 2021. PMID: 34244038 Review.
-
A Review of the Current Practice of Diagnosis and Treatment of Idiopathic Membranous Nephropathy in China.Med Sci Monit. 2021 Feb 7;27:e930097. doi: 10.12659/MSM.930097. Med Sci Monit. 2021. PMID: 33550324 Free PMC article. Review.
References
-
- Rovin BH, Caster DJ, Cattran DC, Gibson KL, Hogan JJ, Moeller MJ, Roccatello D, Cheung M, Wheeler DC, Winkelmayer WC, Floege J; Conference Participants. Management and treatment of glomerular diseases (part 2): conclusions from a Kidney Disease: Improving Global Outcomes (KDIGO) Controversies Conference. Kidney Int. 2019;95:281-95. - PubMed
-
- Keri KC, Blumenthal S, Kulkarni V, Beck L, Chongkrairatanakul T. Primary membranous nephropathy: comprehensive review and historical perspective. Postgrad Med J. 2019;95:23–1. - PubMed
-
- Thokhonelidze I, Maglakelidze N, Sarishvili N, Kasradze T, Dalakishvili K. Association of anti-phospholipasea2-receptor antibodies with clinical course of idiopathic membranous nephropathy. Georgian Med News. 2015;241:49–53. - PubMed
MeSH terms
Substances
Grants and funding
- NO. 20180550523/Liaoning Provincial Natural Science Foundation Project
- NO.2019JH6/10400009/Liaoning Province Central Government's special project to guide local scientific and technological development
- NO.112/3110118034/Major Special Project of Construction Program of China Medical University in 2018
- NO.201902010003/Guangdong Province Major key projects of industrial technology
- NO.50A/The 345 Talent Project of Shengjing Hospital of China Medical University
LinkOut - more resources
Full Text Sources

