Hybrid de novo genome-reassembly reveals new insights on pathways and pathogenicity determinants in rice blast pathogen Magnaporthe oryzae RMg_Dl
- PMID: 34824307
- PMCID: PMC8616942
- DOI: 10.1038/s41598-021-01980-2
Hybrid de novo genome-reassembly reveals new insights on pathways and pathogenicity determinants in rice blast pathogen Magnaporthe oryzae RMg_Dl
Abstract
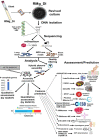
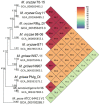
Blast disease incited by Magnaporthe oryzae is a major threat to sustain rice production in all rice growing nations. The pathogen is widely distributed in all rice paddies and displays rapid aerial transmissions, and seed-borne latent infection. In order to understand the genetic variability, host specificity, and molecular basis of the pathogenicity-associated traits, the whole genome of rice infecting Magnaporthe oryzae (Strain RMg_Dl) was sequenced using the Illumina and PacBio (RSII compatible) platforms. The high-throughput hybrid assembly of short and long reads resulted in a total of 375 scaffolds with a genome size of 42.43 Mb. Furthermore, comparative genome analysis revealed 99% average nucleotide identity (ANI) with other oryzae genomes and 83% against M. grisea, and 73% against M. poe genomes. The gene calling identified 10,553 genes with 10,539 protein-coding sequences. Among the detected transposable elements, the LTR/Gypsy and Type LINE showed high occurrence. The InterProScan of predicted protein sequences revealed that 97% protein family (PFAM), 98% superfamily, and 95% CDD were shared among RMg_Dl and reference 70-15 genome, respectively. Additionally, 550 CAZymes with high GH family content/distribution and cell wall degrading enzymes (CWDE) such endoglucanase, beta-glucosidase, and pectate lyase were also deciphered in RMg_Dl. The prevalence of virulence factors determination revealed that 51 different VFs were found in the genome. The biochemical pathway such as starch and sucrose metabolism, mTOR signaling, cAMP signaling, MAPK signaling pathways related genes were identified in the genome. The 49,065 SNPs, 3267 insertions and 3611 deletions were detected, and majority of these varinats were located on downstream and upstream region. Taken together, the generated information will be useful to develop a specific marker for diagnosis, pathogen surveillance and tracking, molecular taxonomy, and species delineation which ultimately leads to device improved management strategies for blast disease.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







References
-
- Sharma TR, et al. Rice blast management through host-plant resistance: retrospect and prospects. Agric. Res. 2012;1:37–52. doi: 10.1007/s40003-011-0003-5. - DOI
-
- Mehta S, Singh B, Dhakate P, Rahman M, Islam MA. Rice, marker-assisted breeding, and disease resistance. In: Wani SH, editor. Disease Resistance in Crop Plants: Molecular, Genetic and Genomic Perspectives. Cham: Springer International Publishing; 2019. pp. 83–111.
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

