Plasmid-Mediated Mechanism of Quinolone Resistance on E. coli Isolates from Different Clinical Samples
- PMID: 34824749
- PMCID: PMC8605851
- DOI: 10.22092/ari.2021.355392.1679
Plasmid-Mediated Mechanism of Quinolone Resistance on E. coli Isolates from Different Clinical Samples
Abstract
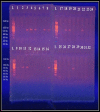
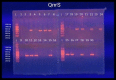
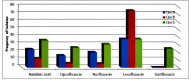
Quinolone antimicrobials are widely used in clinical medicine due to their wide spectrum with high tissue penetration and ease of use; but increasing resistance with clinical use appears to be common in some bacterial pathogens, including Escherichia coli (E.coli). The aim of this study was to investigate plasmid-mediated quinolone resistance determinants (PMQR) including, qnrA, qnrB, and qnrS as the emerging mechanisms of quinolone resistance of E.coli isolates from different clinical sites in Karbala province, Iraq. A total of 200 clinical samples were collected from patients suffering from infections such as UTI, gastro enteritis (diarrhea), vaginitis, and wound infections; 30 samples were diagnosed as E.coli clinical strain from both sexes and different ages after identification by biochemical test, VITEK-2 compact system, and by molecular method using 16Sr DNA marker. Antimicrobial susceptibility and minimal inhibition concentration (MIC) testing for nalidixic acid, norfloxacin, ciprofloxacin, levofloxacin, and gatifloxacin was performed using the broth microdilution method. All strains were screened for PMQR genes qnrA, qnrB, and qnrS by the PCR method after DNA extraction from tested clinical isolates of E.coli. The results showed that E. coli is largely isolated from vaginal (40%) and urine (32%) samples, followed by wound infections (24%) and stools (21%).The high occurrence rate of E. coli(33.33%) isolates was observed in participants aged 31-45 years, while a lower occurrence (10%)was recorded in a group of ˃ 60-year-old female participants. Females have a notably increased frequency of E.coli compared to males, with the female to male ratio being 87%: 13%. Molecular investigation showed the total percentage of E.coli isolates harboring qnr genes to be 21/30 (70%); this figure is composed of 14/30 isolates harboring qnr in combined or mixed form (46.66%) and 7/30 (23.33%) isolates harboring qnr in single form (3 isolates harboring qnrA alone, 1 isolate harboring qnrB alone, 3 isolates harboring qnrS alone).The prevalence rates of qnrA, qnrB, and qnrS were 40%, 43.33%, and 53.33%, respectively. The results also showed that among E.coli isolates encoding qnr genes A, B, and S, 24%, 12%, and 36% were resistant to nalidixic acid, respectively. Among those isolates carrying qnrA, qnrB, and qnrS genes, 15.8%, 5.3%, and 26.3%, respectively, were resistant to ciprofloxacin. Moreover, Norfloxacin resistance was seen in 20.0%, 5.0%, and 30.0% of E.coli isolates harboring qnr A, B, and S genes, respectively. Levofloxacin resistance was seen in 37.5%, 75.0%, and 37.5% of the isolates carrying the qnrA, qnrB, and qnrS genes, respectively. The lowest resistance rates of qnrA, B, and S-positive E.coli strains were against gatifloxacin (0,0, and 25%, respectively).A high prevalence of qnr genes enhances the increasing resistance rate of E.coli against the quinolone antibiotic under study.
Keywords: E.coli; plasmid mediated resistance; qnr; quinolone antibiotics.
Figures


References
-
- Gosling RJ, Clouting CS, Randall LP, Horton RA, Davies RH. Ciprofloxacin resistance in E. coli isolated from turkeys in Great Britain. Avian Pathol. 2012;41(1):83–9. - PubMed
-
- Machuca J, Briales A, Labrador G, Diaz-de-Alba P, Lopez-Rojas R, Docobo-Perez F, et al. Interplay between plasmid-mediated and chromosomal-mediated fluoroquinolone resistance and bacterial fitness in Escherichia coli. J Antimicrob Chemother. 2014;69(12):3203–15. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
