MemDis: Predicting Disordered Regions in Transmembrane Proteins
- PMID: 34830151
- PMCID: PMC8623522
- DOI: 10.3390/ijms222212270
MemDis: Predicting Disordered Regions in Transmembrane Proteins
Abstract
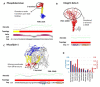
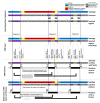
Transmembrane proteins (TMPs) play important roles in cells, ranging from transport processes and cell adhesion to communication. Many of these functions are mediated by intrinsically disordered regions (IDRs), flexible protein segments without a well-defined structure. Although a variety of prediction methods are available for predicting IDRs, their accuracy is very limited on TMPs due to their special physico-chemical properties. We prepared a dataset containing membrane proteins exclusively, using X-ray crystallography data. MemDis is a novel prediction method, utilizing convolutional neural network and long short-term memory networks for predicting disordered regions in TMPs. In addition to attributes commonly used in IDR predictors, we defined several TMP specific features to enhance the accuracy of our method further. MemDis achieved the highest prediction accuracy on TMP-specific dataset among other popular IDR prediction methods.
Keywords: bidirectional long-short term memory; convolutional neural network; deep learning; intrinsically disordered proteins; transmembrane proteins.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

