The Relevance of G-Quadruplexes for DNA Repair
- PMID: 34830478
- PMCID: PMC8620898
- DOI: 10.3390/ijms222212599
The Relevance of G-Quadruplexes for DNA Repair
Abstract
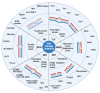
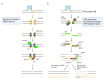
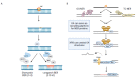
DNA molecules can adopt a variety of alternative structures. Among these structures are G-quadruplex DNA structures (G4s), which support cellular function by affecting transcription, translation, and telomere maintenance. These structures can also induce genome instability by stalling replication, increasing DNA damage, and recombination events. G-quadruplex-driven genome instability is connected to tumorigenesis and other genetic disorders. In recent years, the connection between genome stability, DNA repair and G4 formation was further underlined by the identification of multiple DNA repair proteins and ligands which bind and stabilize said G4 structures to block specific DNA repair pathways. The relevance of G4s for different DNA repair pathways is complex and depends on the repair pathway itself. G4 structures can induce DNA damage and block efficient DNA repair, but they can also support the activity and function of certain repair pathways. In this review, we highlight the roles and consequences of G4 DNA structures for DNA repair initiation, processing, and the efficiency of various DNA repair pathways.
Keywords: G-quadruplex; genome instability; homologous recombination; non-homologous end joining; nucleotide excision repair; translesion synthesis.
Conflict of interest statement
The authors declare no competing interests.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

