Mutations of SARS-CoV-2 spike protein: Implications on immune evasion and vaccine-induced immunity
- PMID: 34836774
- PMCID: PMC8604694
- DOI: 10.1016/j.smim.2021.101533
Mutations of SARS-CoV-2 spike protein: Implications on immune evasion and vaccine-induced immunity
Abstract
Responsible for more than 4.9 million deaths so far, COVID-19, caused by SARS-CoV-2, is instigating devastating effects on the global health care system whose impacts could be longer for the years to come. Acquiring a comprehensive knowledge of host-virus interaction is critical for designing effective vaccines and/or drugs. Understanding the evolution of the virus and the impact of genetic variability on host immune evasion and vaccine efficacy is helpful to design novel strategies to minimize the effects of the emerging variants of concern (VOC). Most vaccines under development and/or in current use target the spike protein owning to its unique function of host receptor binding, relatively conserved nature, potent immunogenicity in inducing neutralizing antibodies, and being a good target of T cell responses. However, emerging SARS-CoV-2 strains are exhibiting variability on the spike protein which could affect the efficacy of vaccines and antibody-based therapies in addition to enhancing viral immune evasion mechanisms. Currently, the degree to which mutations on the spike protein affect immunity and vaccination, and the ability of the current vaccines to confer protection against the emerging variants attracts much attention. This review discusses the implications of SARS-CoV-2 spike protein mutations on immune evasion and vaccine-induced immunity and forward directions which could contribute to future studies focusing on designing effective vaccines and/or immunotherapies to consider viral evolution. Combining vaccines derived from different regions of the spike protein that boost both the humoral and cellular wings of adaptive immunity could be the best options to cope with the emerging VOC.
Keywords: Immunity; Mutation; RBD; SARS-CoV-2; Spike protein; Vaccine.
Copyright © 2021 The Author(s). Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
All authors declare that there is no competing interests.
Figures

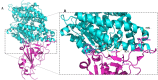

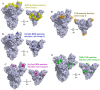

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

