Phylogenetic analysis of the bacterial Pro-Pro-endopeptidase domain reveals a diverse family including secreted and membrane anchored proteins
- PMID: 34841315
- PMCID: PMC8610288
- DOI: 10.1016/j.crmicr.2021.100024
Phylogenetic analysis of the bacterial Pro-Pro-endopeptidase domain reveals a diverse family including secreted and membrane anchored proteins
Abstract
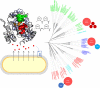
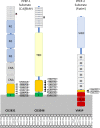
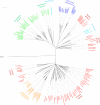
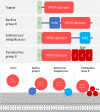
Pro-Pro-endopeptidases (PPEP, EC 3.4.24.89) are secreted, zinc metalloproteases that have the unusual capacity to cleave a peptide bond between two prolines, a bond that is generally less sensitive to proteolytic cleavage. Two well studied members of the family are PPEP-1 and PPEP-2, produced by Clostridioides difficile, a human pathogen, and Paenibacillus alvei, a bee secondary invader, respectively. Both proteases seem to be involved in mediating bacterial adhesion by cleaving cell surface anchor proteins on the bacterium itself. By using basic alignment and phylogenetic profiling analysis, this work shows that the complete family of proteins that contain a PPEP domain includes proteins from more than 130 species spread over 9 genera. These analyses also suggest that the PPEP domain spread through horizontal gene transfer events between species within the Firmicutes' classes Bacilli and Clostridia. Bacterial species containing PPEP homologs are found in diverse habitats, varying from human pathogens and gut microbiota to free-living bacteria, which were isolated from various environments, including extreme conditions such as hot springs, desert soil and salt lakes. The phylogenetic tree reveals the relationships between family members and suggests that smaller subgroups could share cleavage specificity, substrates and functional similarity. Except for PPEP-1 and PPEP-2, no cleavage specificity, specific physiological target, or function has been assigned for any of the other PPEP-family members. Some PPEP proteins have acquired additional domains that recognize and bind noncovalently to various elements of the bacterial peptidoglycan cell-wall, anchoring these PPEPs. Secreted or anchored to the cell-wall surface PPEP proteins seem to perform various functions.
Keywords: Adhesion; Bacterial virulence; CBD, Cell-wall-binding-domain; Cell wall; Endopeptidases; FN3, Fibronectin type III domain; Metalloprotease; Metalloproteases; Motility; MucBP, Mucin binding protein; PPEP, Pro-pro-endopeptidases; Phylogeny; SCWP, Secondary cell-wall polysaccharide; SLH, surface layer homology; Secreted protease; TED, thioester domain; VMSP, VWFA-MucBP-SLH-protein; VWFA, Von Willebrand factor type A.
© 2021 The Authors. Published by Elsevier B.V.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


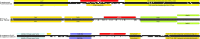
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

