CircEAF2 counteracts Epstein-Barr virus-positive diffuse large B-cell lymphoma progression via miR-BART19-3p/APC/β-catenin axis
- PMID: 34852843
- PMCID: PMC8638185
- DOI: 10.1186/s12943-021-01458-9
CircEAF2 counteracts Epstein-Barr virus-positive diffuse large B-cell lymphoma progression via miR-BART19-3p/APC/β-catenin axis
Abstract
Background: Epstein-Barr virus (EBV) represents an important pathogenic factor of lymphoma and is significantly associated with poor clinical outcome of diffuse large B-cell lymphoma (DLBCL). Circular RNAs (circRNAs) play an essential role in lymphoma progression. However, the underlying mechanism of circRNA on DLBCL progression related to EBV remains largely unknown.
Methods: CircRNA was screened by high-throughput sequencing in tumor samples of 12 patients with DLBCL according to EBV infection status. Expression of circEAF2, as well as the relationship with clinical characteristics and prognosis, were further analyzed in tumor samples of 100 DLBCL patients using quantitative real-time PCR. Gain- and loss-of-function experiments were conducted to investigate the biological functions of circEAF2 both in vitro and in vivo. The underlying mechanism of circRNA on DLBCL progression were further determined by RNA sequencing, RNA pull down assay, dual-luciferase reporter assay, rescue experiments and western blotting.
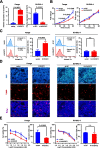
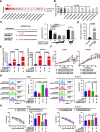
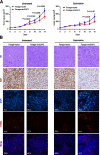
Results: We identified a novel circRNA circEAF2, which was downregulated in EBV + DLBCL and negatively correlated with EBV infection and DLBCL progression. In EBV-positive B lymphoma cells, circEAF2 overexpression induced lymphoma cell apoptosis and sensitized lymphoma cells to epirubicin. As mechanism of action, circEAF2 specifically targeted EBV-encoded miR-BART19-3p, upregulated APC, and suppressed downstream β-catenin expression, resulting in inactivation of Wnt signaling pathway and inhibition of EBV + DLBCL cell proliferation. In EBV-positive B-lymphoma murine models, xenografted tumors with circEAF2 overexpression presented decreased Ki-67 positivity, increased cell apoptosis and retarded tumor growth.
Conclusions: CircEAF2 counteracted EBV + DLBCL progression via miR-BART19-3p/APC/β-catenin axis, referring circEAF2 as a potential prognostic biomarker. Therapeutic targeting EBV-encoded miRNA may be a promising strategy in treating EBV-associated lymphoid malignancies.
Keywords: Diffuse large B-cell lymphoma; Epstein-Barr virus; Wnt signaling pathway; circEAF2; miR-BART19-3p; β-catenin.
© 2021. The Author(s).
Conflict of interest statement
All authors agree to submit for consideration for publication in the journal. All authors declare no conflict of interest.
Figures



Similar articles
-
Epstein-Barr virus-coded miR-BART19-3p promotes proliferation of EBV-associated gastric cancer by inhibiting GADD45B.J Transl Med. 2025 Aug 25;23(1):956. doi: 10.1186/s12967-025-06955-9. J Transl Med. 2025. PMID: 40855561 Free PMC article.
-
Epstein-Barr Virus Infection of Cell Lines Derived from Diffuse Large B-Cell Lymphomas Alters MicroRNA Loading of the Ago2 Complex.J Virol. 2019 Jan 17;93(3):e01297-18. doi: 10.1128/JVI.01297-18. Print 2019 Feb 1. J Virol. 2019. PMID: 30429351 Free PMC article.
-
Lnc SMAD5-AS1 as ceRNA inhibit proliferation of diffuse large B cell lymphoma via Wnt/β-catenin pathway by sponging miR-135b-5p to elevate expression of APC.Cell Death Dis. 2019 Mar 15;10(4):252. doi: 10.1038/s41419-019-1479-3. Cell Death Dis. 2019. PMID: 30874550 Free PMC article.
-
Epstein-Barr virus-positive diffuse large B-cell lymphoma of the elderly: a concise review and update.Curr Opin Oncol. 2015 Sep;27(5):392-8. doi: 10.1097/CCO.0000000000000210. Curr Opin Oncol. 2015. PMID: 26258272 Review.
-
Epstein-Barr virus-positive diffuse large B-cell lymphomas of the elderly.Adv Anat Pathol. 2011 Sep;18(5):349-55. doi: 10.1097/PAP.0b013e318229bf08. Adv Anat Pathol. 2011. PMID: 21841405 Review.
Cited by
-
The Wnt/β-catenin signalling pathway in Haematological Neoplasms.Biomark Res. 2022 Oct 13;10(1):74. doi: 10.1186/s40364-022-00418-9. Biomark Res. 2022. PMID: 36224652 Free PMC article. Review.
-
CircRNAs regulate the crosstalk between inflammation and tumorigenesis: The bilateral association and molecular mechanisms.Mol Ther. 2023 Jun 7;31(6):1514-1532. doi: 10.1016/j.ymthe.2022.12.005. Epub 2022 Dec 14. Mol Ther. 2023. PMID: 36518080 Free PMC article. Review.
-
Circular RNA and Its Roles in the Occurrence, Development, Diagnosis of Cancer.Front Oncol. 2022 Apr 7;12:845703. doi: 10.3389/fonc.2022.845703. eCollection 2022. Front Oncol. 2022. PMID: 35463362 Free PMC article. Review.
-
Mechanism of Action of circRNA/miRNA Network in DLBCL.Noncoding RNA. 2025 Mar 4;11(2):22. doi: 10.3390/ncrna11020022. Noncoding RNA. 2025. PMID: 40126346 Free PMC article. Review.
-
The malignancy of chordomas is enhanced via a circTLK1/miR-16-5p/Smad3 positive feedback axis.Cell Death Discov. 2023 Feb 15;9(1):64. doi: 10.1038/s41420-023-01332-1. Cell Death Discov. 2023. PMID: 36792585 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- 82130004/National Natural Science Foundation of China
- 81830007/National Natural Science Foundation of China
- 81870315/National Natural Science Foundation of China
- 81600155/National Natural Science Foundation of China
- U2005204/National Natural Science Foundation of China
- 19QA145600/Shanghai Rising-Star Program
- 2017YQ075/Municipal Human Resources Development Program for Outstanding Young Talents in Medical and Health Sciences in Shanghai
- 2017XQ2030/Program for Innovative Research Team in Science and technology in Fujian Medical University
- 2019QH2012/Program for Innovative Research Team in Science and technology in Fujian Medical University
- 2016Y9032/Fujian Provincial Department of Science and Technology (CN)
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

