Distinguishing different modes of growth using single-cell data
- PMID: 34854811
- PMCID: PMC8727026
- DOI: 10.7554/eLife.72565
Distinguishing different modes of growth using single-cell data
Abstract
Collection of high-throughput data has become prevalent in biology. Large datasets allow the use of statistical constructs such as binning and linear regression to quantify relationships between variables and hypothesize underlying biological mechanisms based on it. We discuss several such examples in relation to single-cell data and cellular growth. In particular, we show instances where what appears to be ordinary use of these statistical methods leads to incorrect conclusions such as growth being non-exponential as opposed to exponential and vice versa. We propose that the data analysis and its interpretation should be done in the context of a generative model, if possible. In this way, the statistical methods can be validated either analytically or against synthetic data generated via the use of the model, leading to a consistent method for inferring biological mechanisms from data. On applying the validated methods of data analysis to infer cellular growth on our experimental data, we find the growth of length in E. coli to be non-exponential. Our analysis shows that in the later stages of the cell cycle the growth rate is faster than exponential.
Keywords: E. coli; data analysis; infectious disease; linear regression; mathematical model; microbial growth; microbiology; physics of living systems.
Plain language summary
All cells – from bacteria to humans – tightly control their size as they grow and divide. Cells can also change the speed at which they grow, and the pattern of how fast a cell grows with time is called ‘mode of growth’. Mode of growth can be ‘linear’, when cells increase their size at a constant rate, or ‘exponential’, when cells increase their size at a rate proportional to their current size. A cell’s mode of growth influences its inner workings, so identifying how a cell grows can reveal information about how a cell will behave. Scientists can measure the size of cells as they age and identify their mode of growth using single cell imaging techniques. Unfortunately, the statistical methods available to analyze the large amounts of data generated in these experiments can lead to incorrect conclusions. Specifically, Kar et al. found that scientists had been using specific types of plots to analyze growth data that were prone to these errors, and may lead to misinterpreting exponential growth as linear and vice versa. This discrepancy can be resolved by ensuring that the plots used to determine the mode of growth are adequate for this analysis. But how can the adequacy of a plot be tested? One way to do this is to generate synthetic data from a known model, which can have a specific and known mode of growth, and using this data to test the different plots. Kar et al. developed such a ‘generative model’ to produce synthetic data similar to the experimental data, and used these data to determine which plots are best suited to determine growth mode. Once they had validated the best statistical methods for studying mode of growth, Kar et al. applied these methods to growth data from the bacterium Escherichia coli. This showed that these cells have a form of growth called ‘super-exponential growth’. These findings identify a strategy to validate statistical methods used to analyze cell growth data. Furthermore, this strategy – the use of generative models to produce synthetic data to test the accuracy of statistical methods – could be used in other areas of biology to validate statistical approaches.
© 2021, Kar et al.
Conflict of interest statement
PK, ST, JM, JM, AA No competing interests declared
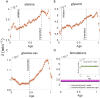
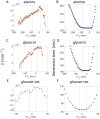
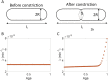
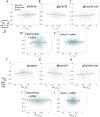
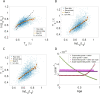
Figures






Similar articles
-
Elongation and surface extension of individual cells of Escherichia coli B/r: comparison of theoretical and experimental size distributions.J Theor Biol. 1987 Dec 7;129(3):337-48. doi: 10.1016/s0022-5193(87)80006-1. J Theor Biol. 1987. PMID: 3332028
-
Biomass growth rate during the prokaryote cell cycle.Crit Rev Microbiol. 1993;19(1):17-42. doi: 10.3109/10408419309113521. Crit Rev Microbiol. 1993. PMID: 8481211 Review.
-
Fission yeast cells grow approximately exponentially.Cell Cycle. 2019 Apr;18(8):869-879. doi: 10.1080/15384101.2019.1595874. Epub 2019 Apr 6. Cell Cycle. 2019. PMID: 30957637 Free PMC article.
-
Volume growth in animal cells is cell cycle dependent and shows additive fluctuations.Elife. 2022 Jan 28;11:e70816. doi: 10.7554/eLife.70816. Elife. 2022. PMID: 35088713 Free PMC article.
-
Growth during the cell cycle.Int Rev Cytol. 2003;226:165-258. doi: 10.1016/s0074-7696(03)01004-0. Int Rev Cytol. 2003. PMID: 12921238 Review.
Cited by
-
Analytical cell size distribution: lineage-population bias and parameter inference.J R Soc Interface. 2022 Nov;19(196):20220405. doi: 10.1098/rsif.2022.0405. Epub 2022 Nov 23. J R Soc Interface. 2022. PMID: 36416039 Free PMC article.
-
Super-exponential growth and stochastic size dynamics in rod-like bacteria.Biophys J. 2023 Apr 4;122(7):1254-1267. doi: 10.1016/j.bpj.2023.02.015. Epub 2023 Feb 22. Biophys J. 2023. PMID: 36814380 Free PMC article.
-
Accurate phenotype-to-genotype mapping of high-diversity yeast libraries by heat-shock-electroporation (HEEL).mBio. 2025 Feb 5;16(2):e0319724. doi: 10.1128/mbio.03197-24. Epub 2024 Dec 20. mBio. 2025. PMID: 39704499 Free PMC article.
-
Gene expression in growing cells: A biophysical primer.ArXiv [Preprint]. 2023 Nov 20:arXiv:2311.12143v1. ArXiv. 2023. PMID: 38045483 Free PMC article. Preprint.
-
Genome concentration limits cell growth and modulates proteome composition in Escherichia coli.Elife. 2024 Dec 23;13:RP97465. doi: 10.7554/eLife.97465. Elife. 2024. PMID: 39714909 Free PMC article.
References
-
- Amir A. Cell Size Regulation in Bacteria. Physical Review Letters. 2014;112:208102. doi: 10.1103/PhysRevLett.112.208102. - DOI
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources

