A Conformationally Stable Acyclic β-Hairpin Scaffold Tolerating the Incorporation of Poorly β-Sheet-Prone Amino Acids
- PMID: 34856053
- PMCID: PMC9299858
- DOI: 10.1002/cbic.202100604
A Conformationally Stable Acyclic β-Hairpin Scaffold Tolerating the Incorporation of Poorly β-Sheet-Prone Amino Acids
Abstract
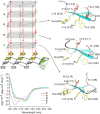
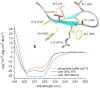
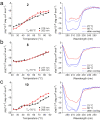
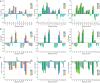
The β-hairpin is a structural element of native proteins, but it is also a useful artificial scaffold for finding lead compounds to convert into peptidomimetics or non-peptide structures for drug discovery. Since linear peptides are synthetically more easily accessible than cyclic ones, but are structurally less well-defined, we propose XWXWXpPXK(/R)X(R) as an acyclic but still rigid β-hairpin scaffold that is robust enough to accommodate different types of side chains, regardless of the secondary-structure propensity of the X residues. The high conformational stability of the scaffold results from tight contacts between cross-strand cationic and aromatic side chains, combined with the strong tendency of the d-Pro-l-Pro dipeptide to induce a type II' β-turn. To demonstrate the robustness of the scaffold, we elucidated the NMR structures and performed molecular dynamics (MD) simulations of a series of peptides displaying mainly non-β-branched, poorly β-sheet-prone residues at the X positions. Both the NMR and MD data confirm that our acyclic β-hairpin scaffold is highly versatile as regards the amino-acid composition of the β-sheet face opposite to the cationic-aromatic one.
Keywords: CH−π interactions; NMR spectroscopy; cation−π interactions; circular dichroism; hairpins.
© 2021 The Authors. ChemBioChem published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



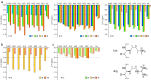

References
-
- Sibanda B. L., Thornton J. M., Nature 1985, 316, 170–174. - PubMed
-
- Iuchi S., in Zinc Finger Proteins: From Atomic Contact to Cellular Function (Eds.: luchi S., Kuldell N.), Landes Bioscience/Eurekah.com and Kluwer Academic/Plenum Publishers, 2005, pp. 7–13.
-
- Blanco F. J., Rivas G., Serrano L., Nat. Struct. Biol. 1994, 1, 584–590. - PubMed
-
- Franklin M. C., Carey K. D., Vajdos F. F., Leahy D. J., de Vos A. M., Sliwkowski M. X., Cancer Cell 2004, 5, 317–328. - PubMed
-
- None
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

