Plant transcription factors - being in the right place with the right company
- PMID: 34856504
- PMCID: PMC8844091
- DOI: 10.1016/j.pbi.2021.102136
Plant transcription factors - being in the right place with the right company
Abstract
Transcriptional regulation underlies many of the growth and developmental processes that shape plants as well as their adaptation to their environment. Key to transcriptional control are transcription factors, DNA-binding proteins that serve two essential functions: to find the appropriate DNA contact sites in their target genes; and to recruit other proteins to execute transcriptional transactions. In recent years, protein structural, genomic, bioinformatic, and proteomic analyses have led to new insights into how these central functions are regulated. Here, we review new findings relating to plant transcription factor function and to their role in shaping transcription in the context of chromatin.
Keywords: Binding specificity; Chromatin; DNA binding; Pioneer transcription factor; Protein condensation; Transcription.
Copyright © 2021 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of competing interest Nothing declared.
Figures
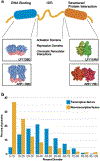

References
-
- Boer DR, Freire-Rios A, van den Berg WAM, Saaki T, Manfield IW, Kepinski S, López-Vidrieo I, Franco-Zorrilla JM, de Vries SC, Solano R, et al. : Structural basis for DNA binding specificity by the auxin-dependent ARF transcription factors. Cell 2014, 156:577–589. - PubMed
-
-
Kato H, Mutte SK, Suzuki H, Crespo I, Das S, Radoeva T, Fontana M, Yoshitake Y, Hainiwa E, van den Berg W, et al. : Design principles of a minimal auxin response system. Nat Plants 2020, 6:473–482.
The authors use the minimal Marchantia auxin response system to derive a simple model for auxin-dependent gene expression, in which auxin-sensitive A- and auxin-insensitive B-class ARFs compete for binding the same target sites in auxin-responsive promoters.
-
-
-
Freire-Rios A, Tanaka K, Crespo I, van der Wijk E, Sizentsova Y, Levitsky V, Lindhoud S, Fontana M, Hohlbein J, Boer DR, et al. : Architecture of DNA elements mediating ARF transcription factor binding and auxin-responsive gene expression in. Proc Natl Acad Sci U S A 2020, 117:24557–24566.
The authors use bioinformatic analysis, coupled to structural biology and Arabidopsis genetics, to demonstrate the requirements for high-affinity DNA binding by ARF transcription factors and rationalize the basis for enrichment of AuxRE repeats in auxin-responsive promoters.
-
-
- Bieluszewski T, Xiao J, Yang Y, Wagner D: PRC2 activity, recruitment, and silencing: a comparative perspective. Trends Plant Sci 2021, 0. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

