Mapping the serum proteome to neurological diseases using whole genome sequencing
- PMID: 34857772
- PMCID: PMC8640022
- DOI: 10.1038/s41467-021-27387-1
Mapping the serum proteome to neurological diseases using whole genome sequencing
Abstract
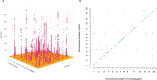
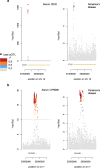
Despite the increasing global burden of neurological disorders, there is a lack of effective diagnostic and therapeutic biomarkers. Proteins are often dysregulated in disease and have a strong genetic component. Here, we carry out a protein quantitative trait locus analysis of 184 neurologically-relevant proteins, using whole genome sequencing data from two isolated population-based cohorts (N = 2893). In doing so, we elucidate the genetic landscape of the circulating proteome and its connection to neurological disorders. We detect 214 independently-associated variants for 107 proteins, the majority of which (76%) are cis-acting, including 114 variants that have not been previously identified. Using two-sample Mendelian randomisation, we identify causal associations between serum CD33 and Alzheimer's disease, GPNMB and Parkinson's disease, and MSR1 and schizophrenia, describing their clinical potential and highlighting drug repurposing opportunities.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
The genetic regulation of protein expression in cerebrospinal fluid.EMBO Mol Med. 2023 Jan 11;15(1):e16359. doi: 10.15252/emmm.202216359. Epub 2022 Dec 12. EMBO Mol Med. 2023. PMID: 36504281 Free PMC article.
-
Identifying drug targets for neurological and psychiatric disease via genetics and the brain transcriptome.PLoS Genet. 2021 Jan 8;17(1):e1009224. doi: 10.1371/journal.pgen.1009224. eCollection 2021 Jan. PLoS Genet. 2021. PMID: 33417599 Free PMC article.
-
Brain Proteome-Wide and Transcriptome-Wide Asso-ciation Studies, Bayesian Colocalization, and Mendelian Randomization Analyses Reveal Causal Genes of Parkinson's Disease.J Gerontol A Biol Sci Med Sci. 2023 Mar 30;78(4):563-568. doi: 10.1093/gerona/glac245. J Gerontol A Biol Sci Med Sci. 2023. PMID: 36511626
-
Microglia in Alzheimer's Disease: Exploring How Genetics and Phenotype Influence Risk.J Mol Biol. 2019 Apr 19;431(9):1805-1817. doi: 10.1016/j.jmb.2019.01.045. Epub 2019 Feb 7. J Mol Biol. 2019. PMID: 30738892 Free PMC article. Review.
-
The age-related microglial transformation in Alzheimer's disease pathogenesis.Neurobiol Aging. 2020 Aug;92:82-91. doi: 10.1016/j.neurobiolaging.2020.03.024. Epub 2020 Apr 15. Neurobiol Aging. 2020. PMID: 32408056 Review.
Cited by
-
Integrated multiple-microarray analysis and mendelian randomization to identify novel targets involved in diabetic nephropathy.Front Endocrinol (Lausanne). 2023 Jul 10;14:1191768. doi: 10.3389/fendo.2023.1191768. eCollection 2023. Front Endocrinol (Lausanne). 2023. PMID: 37492198 Free PMC article.
-
Proteogenomic links to human metabolic diseases.Nat Metab. 2023 Mar;5(3):516-528. doi: 10.1038/s42255-023-00753-7. Epub 2023 Feb 23. Nat Metab. 2023. PMID: 36823471 Free PMC article.
-
Expanding causal genes for Parkinson's disease via multi-omics analysis.NPJ Parkinsons Dis. 2023 Oct 21;9(1):146. doi: 10.1038/s41531-023-00591-0. NPJ Parkinsons Dis. 2023. PMID: 37865667 Free PMC article.
-
Canonical correlation analysis for multi-omics: Application to cross-cohort analysis.PLoS Genet. 2023 May 22;19(5):e1010517. doi: 10.1371/journal.pgen.1010517. eCollection 2023 May. PLoS Genet. 2023. PMID: 37216410 Free PMC article.
-
Blood based biomarkers for movement disorders.Acta Neurol Scand. 2022 Oct;146(4):353-361. doi: 10.1111/ane.13700. Acta Neurol Scand. 2022. PMID: 36156206 Free PMC article. Review.
References
-
- De Jager PL, Yang H-S, Bennett DA. Deconstructing and targeting the genomic architecture of human neurodegeneration. Nat. Neurosci. 2018;21:1310–1317. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

