Genomic and Transcriptomic Analysis of Bovine Pasteurella multocida Serogroup A Strain Reveals Insights Into Virulence Attenuation
- PMID: 34859092
- PMCID: PMC8631534
- DOI: 10.3389/fvets.2021.765495
Genomic and Transcriptomic Analysis of Bovine Pasteurella multocida Serogroup A Strain Reveals Insights Into Virulence Attenuation
Abstract
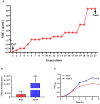
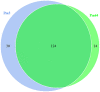
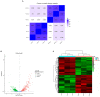
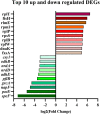
Pasteurella multocida is one of the primary pathogens of bovine respiratory disease (BRD), and causes huge losses in the cattle industry. The Pm3 strain was a natural isolate, which is a strong form of pathogen and is sensitive to fluoroquinolones antibiotics. A high fluoroquinolone resistant strain, Pm64 (MIC = 64 μg/mL), was formed after continuous induction with subinhibitory concentration (1/2 MIC) of enrofloxacin, with the enhanced growth characteristics and large attenuation of pathogenicity in mice. This study reports the whole genome sequence and the transcription profile by RNA-Seq of strain Pm3/Pm64. The results showed an ineffective difference between the two strains at the genome level. However, 32 genes could be recognized in the gene islands (GIs) of Pm64, in which 24 genes were added and 8 genes were lost. Those genes are involved in DNA binding, trehalose metabolism, material transportation, capsule synthesis, prophage, amino acid metabolism, and other functions. In Pm3 strain, 558 up-regulated and 568 down-regulated genes were found compared to Pm64 strain, from which 20 virulence factor-related differentially expressed genes (DEGs) were screened. Mainly differentially transcribed genes were associated with capsular polysaccharide (CPS), lipopolysaccharide (LPS), lipooligosaccharide (LOS). Iron utilization, and biofilm composition. We speculated that the main mechanism of virulence attenuation after the formation of resistance of Pm64 comes from the change of the expression profile of these genes. This report elucidated the toxicity targets of P. multocida serogroup A which provide fundamental information toward the understanding of the pathogenic mechanism and to decreasing antimicrobial drugs resistance.
Keywords: P. multocida serogroup A; complete genome sequencing; gene islands; transcriptomic sequencing; virulence gene.
Copyright © 2021 Zhan, Zhang, Zhao, Li, Zhang, Hu, Elken, Kong and Gao.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Capsular Polysaccharide Interferes with Biofilm Formation by Pasteurella multocida Serogroup A.mBio. 2017 Nov 21;8(6):e01843-17. doi: 10.1128/mBio.01843-17. mBio. 2017. PMID: 29162713 Free PMC article.
-
Transcriptomic Analysis of High- and Low-Virulence Bovine Pasteurella multocida in vitro and in vivo.Front Vet Sci. 2021 Feb 10;8:616774. doi: 10.3389/fvets.2021.616774. eCollection 2021. Front Vet Sci. 2021. PMID: 33644147 Free PMC article.
-
Experimental pathogenicity and complete genome characterization of a pig origin Pasteurella multocida serogroup F isolate HN07.Vet Microbiol. 2017 Jan;198:23-33. doi: 10.1016/j.vetmic.2016.11.028. Epub 2016 Nov 27. Vet Microbiol. 2017. PMID: 28062004
-
Pasteurella multocida and bovine respiratory disease.Anim Health Res Rev. 2007 Dec;8(2):129-50. doi: 10.1017/S1466252307001399. Anim Health Res Rev. 2007. PMID: 18218157 Review.
-
Candidate vaccine antigens and genes in Pasteurella multocida.J Biotechnol. 1999 Aug 20;73(2-3):83-90. doi: 10.1016/s0168-1656(99)00111-x. J Biotechnol. 1999. PMID: 10486918 Review.
Cited by
-
The Research Status, Potential Hazards and Toxicological Mechanisms of Fluoroquinolone Antibiotics in the Environment.Antibiotics (Basel). 2023 Jun 15;12(6):1058. doi: 10.3390/antibiotics12061058. Antibiotics (Basel). 2023. PMID: 37370377 Free PMC article. Review.
-
Role of biofilms in antimicrobial resistance of the bacterial bovine respiratory disease complex.Front Vet Sci. 2024 Jun 12;11:1353551. doi: 10.3389/fvets.2024.1353551. eCollection 2024. Front Vet Sci. 2024. PMID: 38933702 Free PMC article. Review.
-
Integration of Non-Coding RNA and mRNA Profiles Reveals the Mechanisms of Rumen Development Induced by Different Types of Diet in Calves.Genes (Basel). 2023 May 16;14(5):1093. doi: 10.3390/genes14051093. Genes (Basel). 2023. PMID: 37239453 Free PMC article.
-
Transcriptomic Changes and satP Gene Function Analysis in Pasteurella multocida with Different Levels of Resistance to Enrofloxacin.Vet Sci. 2023 Mar 28;10(4):257. doi: 10.3390/vetsci10040257. Vet Sci. 2023. PMID: 37104412 Free PMC article.
-
Genomic and transcriptomics analysis reveal putative secreted proteins expressed of Pasteurella multocida during 18β-glycyrrhetinic acid treatment.Front Vet Sci. 2024 Nov 7;11:1495924. doi: 10.3389/fvets.2024.1495924. eCollection 2024. Front Vet Sci. 2024. PMID: 39575438 Free PMC article.
References
LinkOut - more resources
Full Text Sources

