Structure, Function and Regulation of a Second Pyruvate Kinase Isozyme in Pseudomonas aeruginosa
- PMID: 34867929
- PMCID: PMC8637920
- DOI: 10.3389/fmicb.2021.790742
Structure, Function and Regulation of a Second Pyruvate Kinase Isozyme in Pseudomonas aeruginosa
Abstract
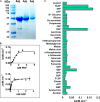
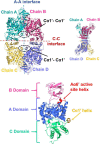
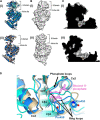
Pseudomonas aeruginosa (PA) depends on the Entner-Doudoroff pathway (EDP) for glycolysis. The main enzymatic regulator in the lower half of the EDP is pyruvate kinase. PA contains genes that encode two isoforms of pyruvate kinase, denoted PykAPA and PykFPA. In other well-characterized organisms containing two pyruvate kinase isoforms (such as Escherichia coli) each isozyme is differentially regulated. The structure, function and regulation of PykAPA has been previously characterized in detail, so in this work, we set out to assess the biochemical and structural properties of the PykFPA isozyme. We show that pykF PA expression is induced in the presence of the diureide, allantoin. In spite of their relatively low amino acid sequence identity, PykAPA and PykFPA display broadly comparable kinetic parameters, and are allosterically regulated by a very similar set of metabolites. However, the x-ray crystal structure of PykFPA revealed significant differences compared with PykAPA. Notably, although the main allosteric regulator binding-site of PykFPA was empty, the "ring loop" covering the site adopted a partially closed conformation. Site-directed mutation of the proline residues flanking the ring loop yielded apparent "locked on" and "locked off" allosteric activation phenotypes, depending on the residue mutated. Analysis of PykFPA inter-protomer interactions supports a model in which the conformational transition(s) accompanying allosteric activation involve re-orientation of the A and B domains of the enzyme and subsequent closure of the active site.
Keywords: Entner-Doudoroff pathway; Pseudomonas aeruginosa; bacterial metabolism; glycolysis; pykF; pyruvate kinase; x-ray crystallography.
Copyright © 2021 Abdelhamid, Wang, Parkhill, Brear, Chee, Rahman and Welch.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



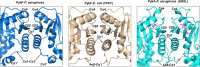
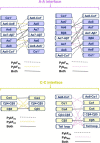
References
-
- Al-Zaid Siddiquee K., Arauzo-Bravo M. J., Shimizu K. (2004). Metabolic flux analysis of pykF gene knockout Escherichia coli based on 13C-labeling experiments together with measurements of enzyme activities and intracellular metabolite concentrations. Appl. Microbiol. Biotechnol. 63 407–417. 10.1007/s00253-003-1357-9 - DOI - PubMed
LinkOut - more resources
Full Text Sources

