Phylogenetic Analysis of the Plant U2 snRNP Auxiliary Factor Large Subunit A Gene Family in Response to Developmental Cues and Environmental Stimuli
- PMID: 34868124
- PMCID: PMC8635922
- DOI: 10.3389/fpls.2021.739671
Phylogenetic Analysis of the Plant U2 snRNP Auxiliary Factor Large Subunit A Gene Family in Response to Developmental Cues and Environmental Stimuli
Abstract
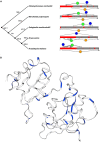
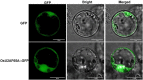
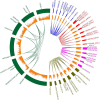
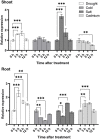
In all organisms, splicing occurs through the formation of spliceosome complexes, and splicing auxiliary factors are essential during splicing. U2AF65 is a crucial splicing cofactor, and the two typical RNA-recognition motifs at its center recognize and bind the polypyrimidine sequence located between the intron branch site and the 3'-splice site. U2AF65A is a member of the U2AF65 gene family, with pivotal roles in diseases in mammals, specifically humans; however, few studies have investigated plant U2AF65A, and its specific functions are poorly understood. Therefore, in the present study, we systematically identified U2AF65A in plant species from algae to angiosperms. Based on 113 putative U2AF65A sequences from 33 plant species, phylogenetic analyses were performed, followed by basic bioinformatics, including the comparisons of gene structure, protein domains, promoter motifs, and gene expression levels. In addition, using rice as the model crop, we demonstrated that the OsU2AF65A protein is localized to the nucleus and cytoplasm, and it is involved in responses to various stresses, such as drought, high salinity, low temperature, and heavy metal exposure (e.g., cadmium). Using Arabidopsis thaliana and rice mutants, we demonstrated that U2AF65A is involved in the accumulation of plant biomass, growth of hypocotyl upon thermal stimulation, and reduction of tolerance of high temperature stress. These findings offer an overview of the U2AF65 gene family and its stress response functions, serving as the reference for further comprehensive functional studies of the essential specific splicing cofactor U2AF65A in the plant kingdom.
Keywords: U2AF65A; gene expression; protein interaction; splicing; stress.
Copyright © 2021 Lu, Gao, Wang, He, Du, Chen, Zhao, Fang, Wang and Cao.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Regulation of flowering transition by alternative splicing: the role of the U2 auxiliary factor.J Exp Bot. 2020 Jan 23;71(3):751-758. doi: 10.1093/jxb/erz416. J Exp Bot. 2020. PMID: 31605606
-
Phylogenetic comparison of 5' splice site determination in central spliceosomal proteins of the U1-70K gene family, in response to developmental cues and stress conditions.Plant J. 2020 Jul;103(1):357-378. doi: 10.1111/tpj.14735. Epub 2020 Apr 24. Plant J. 2020. PMID: 32133712
-
Mammalian splicing factor SF1 interacts with SURP domains of U2 snRNP-associated proteins.Nucleic Acids Res. 2015 Dec 2;43(21):10456-73. doi: 10.1093/nar/gkv952. Epub 2015 Sep 29. Nucleic Acids Res. 2015. PMID: 26420826 Free PMC article.
-
How Are Short Exons Flanked by Long Introns Defined and Committed to Splicing?Trends Genet. 2016 Oct;32(10):596-606. doi: 10.1016/j.tig.2016.07.003. Epub 2016 Aug 6. Trends Genet. 2016. PMID: 27507607 Review.
-
Cancer-Associated Perturbations in Alternative Pre-messenger RNA Splicing.Cancer Treat Res. 2013;158:41-94. doi: 10.1007/978-3-642-31659-3_3. Cancer Treat Res. 2013. PMID: 24222354 Review.
Cited by
-
Differential alternative splicing genes and isoform co-expression networks of Brassica napus under multiple abiotic stresses.Front Plant Sci. 2022 Oct 13;13:1009998. doi: 10.3389/fpls.2022.1009998. eCollection 2022. Front Plant Sci. 2022. PMID: 36311064 Free PMC article.
-
The OsCBL8-OsCIPK17 Module Regulates Seedling Growth and Confers Resistance to Heat and Drought in Rice.Int J Mol Sci. 2022 Oct 18;23(20):12451. doi: 10.3390/ijms232012451. Int J Mol Sci. 2022. PMID: 36293306 Free PMC article.
-
Phylogenetic analysis and stress response of the plant U2 small nuclear ribonucleoprotein B″ gene family.BMC Genomics. 2022 Nov 8;23(1):744. doi: 10.1186/s12864-022-08956-0. BMC Genomics. 2022. PMID: 36348279 Free PMC article.
-
RNA-Binding Protein-Mediated Alternative Splicing Regulates Abiotic Stress Responses in Plants.Int J Mol Sci. 2024 Sep 30;25(19):10548. doi: 10.3390/ijms251910548. Int J Mol Sci. 2024. PMID: 39408875 Free PMC article. Review.
-
The splicing auxiliary factor OsU2AF35a enhances thermotolerance via protein separation and promoting proper splicing of OsHSA32 pre-mRNA in rice.Plant Biotechnol J. 2025 Apr;23(4):1308-1328. doi: 10.1111/pbi.14587. Epub 2025 Jan 22. Plant Biotechnol J. 2025. PMID: 39844526 Free PMC article.
References
LinkOut - more resources
Full Text Sources

