Weighted Gene Co-Expression Network Analysis and Treatment Strategies of Tumor Recurrence-Associated Hub Genes in Lung Adenocarcinoma
- PMID: 34868230
- PMCID: PMC8636777
- DOI: 10.3389/fgene.2021.756235
Weighted Gene Co-Expression Network Analysis and Treatment Strategies of Tumor Recurrence-Associated Hub Genes in Lung Adenocarcinoma
Abstract
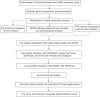
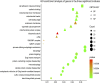
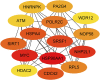
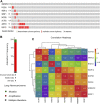
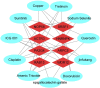
Despite the recent progress of lung adenocarcinoma (LUAD) therapy, tumor recurrence remained to be a challenging factor that impedes the effectiveness of treatment. The objective of the present study was to predict the hub genes affecting LUAD recurrence via weighted gene co-expression network analysis (WGCNA). Microarray samples from LUAD dataset of GSE32863 were analyzed, and the modules with the highest correlation to tumor recurrence were selected. Functional enrichment analysis was conducted, followed by establishment of a protein-protein interaction (PPI) network. Subsequently, hub genes were identified by overall survival analyses and further validated by evaluation of expression in both myeloid populations and tissue samples of LUAD. Gene set enrichment analysis (GSEA) was then carried out, and construction of transcription factors (TF)-hub gene and drug-hub gene interaction network was also achieved. A total of eight hub genes (ACTR3, ARPC5, RAB13, HNRNPK, PA2G4, WDR12, SRSF1, and NOP58) were finally identified to be closely correlated with LUAD recurrence. In addition, TFs that regulate hub genes have been predicted, including MYC, PML, and YY1. Finally, drugs including arsenic trioxide, cisplatin, Jinfukang, and sunitinib were mined for the treatment of the eight hub genes. In conclusion, our study may facilitate the invention of targeted therapeutic drugs and shed light on the understanding of the mechanism for LUAD recurrence.
Keywords: hub genes; lung adenocarcinoma; transcription factor; tumor recurrence; weighted gene co-expression network analysis.
Copyright © 2021 Shen, Liu, Liu, Liu and Yao.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures






References
LinkOut - more resources
Full Text Sources
Miscellaneous

