Molecular Strategies to Target Protein Aggregation in Huntington's Disease
- PMID: 34869596
- PMCID: PMC8636123
- DOI: 10.3389/fmolb.2021.769184
Molecular Strategies to Target Protein Aggregation in Huntington's Disease
Abstract
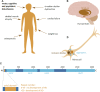
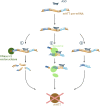
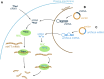
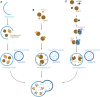
Huntington's disease (HD) is a neurodegenerative disorder caused by the aggregation of the mutant huntingtin (mHTT) protein in nerve cells. mHTT self-aggregates to form soluble oligomers and insoluble fibrils, which interfere in a number of key cellular functions. This leads to cell quiescence and ultimately cell death. There are currently still no treatments available for HD, but approaches targeting the HTT levels offer systematic, mechanism-driven routes towards curing HD and other neurodegenerative diseases. This review summarizes the current state of knowledge of the mRNA targeting approaches such as antisense oligonucleotides and RNAi system; and the novel methods targeting mHTT and aggregates for degradation via the ubiquitin proteasome or the autophagy-lysosomal systems. These methods include the proteolysis-targeting chimera, Trim-Away, autophagosome-tethering compound, autophagy-targeting chimera, lysosome-targeting chimera and approach targeting mHTT for chaperone-mediated autophagy. These molecular strategies provide a knowledge-based approach to target HD and other neurodegenerative diseases at the origin.
Keywords: aggregation; huntingtin (HTT); huntington’s disease; mRNA degradation; protein degradation; protein fibrils; protein quality control; proteostasis.
Copyright © 2021 Jarosińska and Rüdiger.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Ubiquitin-modifying enzymes in Huntington's disease.Front Mol Biosci. 2023 Feb 8;10:1107323. doi: 10.3389/fmolb.2023.1107323. eCollection 2023. Front Mol Biosci. 2023. PMID: 36926679 Free PMC article. Review.
-
Role of TFEB in Huntington's Disease.Biology (Basel). 2024 Apr 4;13(4):238. doi: 10.3390/biology13040238. Biology (Basel). 2024. PMID: 38666850 Free PMC article. Review.
-
Siah-1-interacting protein regulates mutated huntingtin protein aggregation in Huntington's disease models.Cell Biosci. 2022 Mar 19;12(1):34. doi: 10.1186/s13578-022-00755-0. Cell Biosci. 2022. PMID: 35305696 Free PMC article.
-
Proteostasis of Huntingtin in Health and Disease.Int J Mol Sci. 2017 Jul 19;18(7):1568. doi: 10.3390/ijms18071568. Int J Mol Sci. 2017. PMID: 28753941 Free PMC article. Review.
-
Herp Promotes Degradation of Mutant Huntingtin: Involvement of the Proteasome and Molecular Chaperones.Mol Neurobiol. 2018 Oct;55(10):7652-7668. doi: 10.1007/s12035-018-0900-8. Epub 2018 Feb 12. Mol Neurobiol. 2018. PMID: 29430620
Cited by
-
Advances in Stem Cell Therapy for Huntington's Disease: A Comprehensive Literature Review.Cells. 2025 Jan 3;14(1):42. doi: 10.3390/cells14010042. Cells. 2025. PMID: 39791743 Free PMC article.
-
The Other Side of Plastics: Bioplastic-Based Nanoparticles for Drug Delivery Systems in the Brain.Pharmaceutics. 2023 Oct 28;15(11):2549. doi: 10.3390/pharmaceutics15112549. Pharmaceutics. 2023. PMID: 38004530 Free PMC article. Review.
-
The role of Poly-ADP ribose polymerase (PARP) enzymes in chemotherapy-induced cognitive impairments - parallels with other neurodegenerative disorders.Front Pharmacol. 2025 Jun 9;16:1615843. doi: 10.3389/fphar.2025.1615843. eCollection 2025. Front Pharmacol. 2025. PMID: 40552150 Free PMC article. Review.
-
Decoding Neurodegeneration: A Review of Molecular Mechanisms and Therapeutic Advances in Alzheimer's, Parkinson's, and ALS.Int J Mol Sci. 2024 Nov 24;25(23):12613. doi: 10.3390/ijms252312613. Int J Mol Sci. 2024. PMID: 39684324 Free PMC article. Review.
-
Luteolin as potential treatment for Huntington's disease: Insights from a transgenic mouse model.CNS Neurosci Ther. 2024 Sep;30(9):e70025. doi: 10.1111/cns.70025. CNS Neurosci Ther. 2024. PMID: 39228080 Free PMC article.
References
-
- Alterman J. F., Godinho B. M. D. C., Hassler M. R., Ferguson C. M., Echeverria D., Sapp E., et al. (2019). A Divalent siRNA Chemical Scaffold for Potent and Sustained Modulation of Gene Expression throughout the central Nervous System. Nat. Biotechnol. 37 (August), 884–894. 10.1038/s41587-019-0205-0 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources

