Epidemiological Analysis From 2018 to 2020 in China and Prevention Strategy of Porcine Circovirus Type 2
- PMID: 34869727
- PMCID: PMC8635018
- DOI: 10.3389/fvets.2021.753297
Epidemiological Analysis From 2018 to 2020 in China and Prevention Strategy of Porcine Circovirus Type 2
Abstract
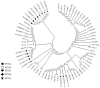
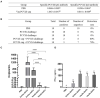
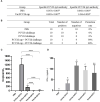
Porcine circovirus type 2 (PCV2) is one of the smallest known animal viruses and is the main pathogen of PCV-associated diseases (PCVAD). Epidemiological surveillance results have shown that the PCV2 infection rate is on the rise in China, thus, PCV2 disease prevention and control has become a huge challenge for the Chinese swine industry. We collected clinical samples from multiple different provinces in China from 2018 to 2020 and found that the positive rate of PCV2 was 53% (3619/6872), identity between the cloned 62 ORF2 genes was 84.4-100% and identity between the cloned 62 ORF2 sequences and reference sequence was 72.9-99.8%. Genetic evolution analysis found that PCV2d accounted for 79% (49/62 samples), PCV2a for 12.9% (8/62 samples), PCV2b for 8% (5/62 samples), and PCV2c and PCV2e genotypes were not found. However, most commercial PCV2 subunit vaccines are based on the PCV2a genotype, and there are very few vaccines based on PCV2b or PCV2d. Therefore, the homologous and heterologous protection ability of PCV2b and PCV2d Cap proteins based on the baculovirus against the PCV2b and PCV2d infections was evaluated, which is expected to design and develop excellent PCV2 protein vaccine candidates. This study found that both PCV2b and PCV2d Cap proteins can increase the level of humoral immunity and cellular immune response in mice. Importantly, both PCV2b and PCV2d cap proteins can provide homologous and heterologous protection against the PCV2b and PCV2d viruses. Overall, this study provides a reference for the prevention and control of PCVAD in mainland China and the development of PCV2 vaccines.
Keywords: PCV2b Cap; PCV2d Cap; epidemiological analysis; heterologous protection; homologous protection.
Copyright © 2021 Huang, Chen, Long, Yang, Song, Liu, Li, Liang, Yu, Huang and Tang.
Conflict of interest statement
All authors are employed by Wuhan Keqian Biology Co., Ltd.
Figures




References
LinkOut - more resources
Full Text Sources
Miscellaneous

