A clinical laboratory-developed LSC17 stemness score assay for rapid risk assessment of patients with acute myeloid leukemia
- PMID: 34872104
- PMCID: PMC8945314
- DOI: 10.1182/bloodadvances.2021005741
A clinical laboratory-developed LSC17 stemness score assay for rapid risk assessment of patients with acute myeloid leukemia
Abstract
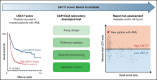
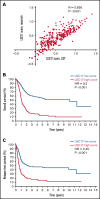
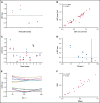
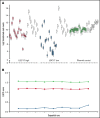
Leukemia stem cells (LSCs) are linked to relapse in acute myeloid leukemia (AML). The LSC17 gene expression score robustly captures LSC stemness properties in AML and can be used to predict survival outcomes and response to therapy, enabling risk-adapted, upfront treatment approaches. The LSC17 score was developed and validated in a research setting. To enable widespread use of the LSC17 score in clinical decision making, we established a laboratory-developed test (LDT) for the LSC17 score that can be deployed broadly in clinical molecular diagnostic laboratories. We extensively validated the LSC17 LDT in a College of American Pathologists/Clinical Laboratory Improvements Act (CAP/CLIA)-certified laboratory, determining specimen requirements, a synthetic control, and performance parameters for the assay. Importantly, we correlated values from the LSC17 LDT to clinical outcome in a reference cohort of patients with AML, establishing a median assay value that can be used for clinical risk stratification of individual patients with newly diagnosed AML. The assay was established in a second independent CAP/CLIA-certified laboratory, and its technical performance was validated using an independent cohort of patient samples, demonstrating that the LSC17 LDT can be readily implemented in other settings. This study enables the clinical use of the LSC17 score for upfront risk-adapted management of patients with AML.
© 2022 by The American Society of Hematology. Licensed under Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International (CC BY-NC-ND 4.0), permitting only noncommercial, nonderivative use with attribution. All other rights reserved.
Figures

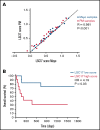
References
-
- Arber DA, Orazi A, Hasserjian R, et al. . The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127(20):2391-2405. - PubMed
-
- Ng SW, Mitchell A, Kennedy JA, et al. . A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature. 2016;540(7633): 433-437. - PubMed
-
- Duployez N, Marceau-Renaut A, Villenet C, et al. . The stem cell-associated gene expression signature allows risk stratification in pediatric acute myeloid leukemia. Leukemia. 2019;33(2):348-357. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

