The role of HIRA-dependent H3.3 deposition and its modifications in the somatic hypermutation of immunoglobulin variable regions
- PMID: 34873043
- PMCID: PMC8685899
- DOI: 10.1073/pnas.2114743118
The role of HIRA-dependent H3.3 deposition and its modifications in the somatic hypermutation of immunoglobulin variable regions
Abstract
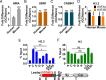
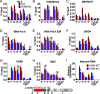
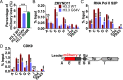
The H3.3 histone variant and its chaperone HIRA are involved in active transcription, but their detailed roles in regulating somatic hypermutation (SHM) of immunoglobulin variable regions in human B cells are not yet fully understood. In this study, we show that the knockout (KO) of HIRA significantly decreased SHM and changed the mutation pattern of the variable region of the immunoglobulin heavy chain (IgH) in the human Ramos B cell line without changing the levels of activation-induced deaminase and other major proteins known to be involved in SHM. Except for H3K79me2/3 and Spt5, many factors related to active transcription, including H3.3, were substantively decreased in HIRA KO cells, and this was accompanied by decreased nascent transcription in the IgH locus. The abundance of ZMYND11 that specifically binds to H3.3K36me3 on the IgH locus was also reduced in the HIRA KO. Somewhat surprisingly, HIRA loss increased the chromatin accessibility of the IgH V region locus. Furthermore, stable expression of ectopic H3.3G34V and H3.3G34R mutants that inhibit both the trimethylation of H3.3K36 and the recruitment of ZMYND11 significantly reduced SHM in Ramos cells, while the H3.3K79M did not. Consistent with the HIRA KO, the H3.3G34V mutant also decreased the occupancy of various elongation factors and of ZMYND11 on the IgH variable and downstream switching regions. Our results reveal an unrecognized role of HIRA and the H3.3K36me3 modification in SHM and extend our knowledge of how transcription-associated chromatin structure and accessibility contribute to SHM in human B cells.
Keywords: HIRA and H3.3 modifications; chromatin accessibility; immunoglobulin variable region; somatic hypermutation; transcription elongation.
Conflict of interest statement
The authors declare no competing interest.
Figures


References
-
- Rogozin I. B., Diaz M., Cutting edge: DGYW/WRCH is a better predictor of mutability at G:C bases in Ig hypermutation than the widely accepted RGYW/WRCY motif and probably reflects a two-step activation-induced cytidine deaminase-triggered process. J. Immunol. 172, 3382–3384 (2004). - PubMed
-
- de Yébenes V. G., Ramiro A. R., Activation-induced deaminase: Light and dark sides. Trends Mol. Med. 12, 432–439 (2006). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

